BAFRegress
bafRegress is a software that detects and estimates sample contamination using B allele frequency data from Illumina genotyping arrays using a regression model.
Download bafRegress
- File:BafRegress.tar.gz (version 0.9.3, 2013-07-30)
This package should contain bafRegress.py and bafRegress.R
About bafRegress
bafRegress is actually a pair of scripts, one Python (2.7) and one R. The Python script is the main interface and handles the parsing of Illumina final report files. The R script actually performs the regression model and provides the contamination estimates.
To use this program, you will need Final Report files from BeadStudio/GenomeStudio. You will need to have columns for the genotypes using the A/B Illumina coded alleles as well as the Illumina reported B allele frequency in addition to the standard sample/marker columns.
Additionally, you will also need allele frequency estimates for the markers on the genotype array. The file should have two columns: one with a marker name that matches the name in the Final Report file and another with the allele frequency for one of the two alleles with a value between 0.0-1.0 or NA if missing.
Basic Usage
Processing Raw Final Report Files
If you have a final report file with exactly one sample in it, the most basic usage of the program is
$ python bafRegress.py estimate --freqfile popmaf.txt finalreport.txt > results.txt
The first parameter "estimate" tells the script you would like to estimate the level of contamination for the sample. You may specify as many Final Report files as you would like on the command line. By default, only the first sample in the file will be analyzed. The results are streamed to stdout, so be sure to redirect the output to a file.
If you have more than one sample per Final Report file, add the "--stacked" option
$ python bafRegress.py estimate --freqfile popmaf.txt --stacked finalreport.txt > results.txt
Processing Binary Files
Since Final Report files can be quite large and much of the work in the scripts is spend parsing the text files and only sending what is necessary to R to minimize memory usage. It can also be somewhat time consuming to repeatedly merge in the allele frequency data. Therefore we've also include an option to create binary data files which can be loaded and accessed much more quickly. This is especially true when there are multiple samples per file.
To convert a standard Final Report to a binary format, use
$ python bafRegress.py convert --outprefix mydata --freqfile popmaf.txt finalreport.txt
Here, a "--freqfile" is not required, but if provided, it will make merging the values during analysis much quicker. You may specify multiple Final Report files as arguments if you would like to merge them into the same binary format. The binary files will all start with the prefix specified by "--outprefix". Once created, you can test all the samples with
$ python bafRegress.py estimatebin mydata > results.txt
If you did not provide a "--freqfile" before, you may specify one at this step.
Final Report Options
Each of the actions "estimate","estimatebin", and "convert" also accept options for the different column names in the Final Report files. If your names to not match the expected names, you may override them. The options are
- --colsample=col, sample column [default: Sample Name]
- --colmarker=col, marker column [default: SNP Name]
- --colbaf=col, B allele frequency column [default: B Allele Freq]
- --colab1=col, AB Allele 1 column [default: Allele1 - AB]
- --colab2=col, AB Allele 2 column [default: Allele2 - AB]
Subset Options
If you would only like to include certain markers (or exclude certain markers) there are options to do that as well. (Note, when using binary files, you will only need to do this during the "convert" step.). These options happen to follow the same parameter names used by PLINK.
- --extract=FILE Include only these markers
- --exclude=FILE Include all but these markers
- --keep=FILE Include only these individuals
- --remove=FILE Include all but these individuals
By default these files are split by white space and the first column is used. The sample name or marker name must match exactly to what is in the Final Report. If you want to change the field separator or the column used for option "X", you may set "--Xcol" or "--Xsep" respectively (eg --extractsep="," or --keepcol=2)
Frequency File Options
A file with frequency information is needed for analsis. You can specify
- --freqfile=FILE, Text file with marker name and population frequency
- --freqcols=COLS, Columns for marker/frequency [default 1,2]
- --freqsep=SEP, Split FILE using this [default: white space]
You must specify two columns, the marker and the frequency (in that order). So if you use PLINK to calculate allele frequencies, you would specify "--freqcols=2,5". Also note that only the minor allele frequency is used, so if the frequency provided is >0.50, it is automatically reduced by 0.50 during analysis.
Plotting
We're provided basic support for plotting samples if you would like to visually inspect them for anomalies as well. You would use
$ python bafRegress.py plot --freqfile popmaf.txt finalreport.txt
or
$ python bafRegress.py plotbin --sample SAMPLENAME mydata
depending on whether you are using the Final Report files or the binary versions.
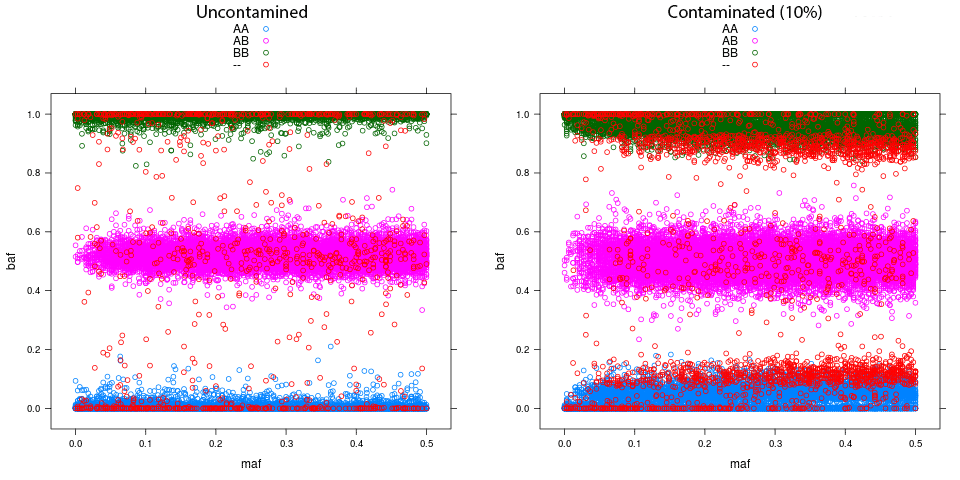
Here is an example of the plot for an uncontaminated vs a contaminated sample. Note how the uncontaminated sample has values closer to the expected BAF of 0 and 1 for the homozygotes. Also note that in the contaminated sample, the deviance from the expected seems to increase as a function of the minor allele frequency (MAF).
Interpreting Results
For each sample, the following values are returned:
- estimate - This is the estimate of contamination from the model (on the 0.0-1.0 scale)
- stderr - The standard error of the estimate
- tval - The test statistic
- pval - The p-value of the estimate
- callrate - The call rate of the sample (number of non-missing genotypes)
- Nhom - The number of homozygous genotypes used to fit the model
You would typically set an estimate cutoff of 1%-2% as a lower bound. If you have many SNPs, sometimes the result can have a significant p-value but the detected level of contamination is like 0.2% which is practically insignificant. This regression method is effective at quantifying modest levels of contamination. More extreme levels of contamination are harder to estimate, but will have a dramatic effect on the call rate making it go way down. You'll probably want to set a minimum reasonable call rate threshold on the samples as well (say, 95%).
Finally, It is useful to look at Nhom to see how many markers were used to create the estimate. You'll want to make sure you're using at least 1,000 markers to get good estimates. With a typical genome-wide array, this shouldn't be a problem at all.
Reference
Please cite the following paper:
G. Jun, M. Flickinger, K. N. Hetrick, Kurt, J. M. Romm, K. F. Doheny, G. Abecasis, M. Boehnke,and H. M. Kang, Detecting and Estimating Contamination of Human DNA Samples in Sequencing and Array-Based Genotype Data, American journal of human genetics doi:10.1016/j.ajhg.2012.09.004 (volume 91 issue 5 pp.839 - 848)