GotCloud: Alignment Pipeline
Back to parent: GotCloud
The Mapping Pipeline takes FASTQ files and generates recalibrated BAM files from them.
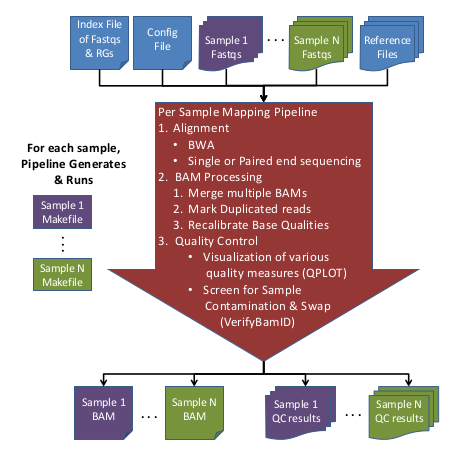
Here is an overview of the Mapping Pipeline:
Input Data:
- Raw Sequence (FASTQ) files
- Sequence Index file containing fastqs & RG info
- Reference files
- (Optional) Configuration file to override default options
Raw Sequence (FASTQ) files
These are the FASTQ files that need to be mapped to BAM files.
These files are specified in the Sequence Index File.
Sequence Index File
This file specifies the FASTQ files that need to be processed and the Read Group information for them.
This file is specified either via the command line parameter --index_file or via the configuration file setting INDEX_FILE.
The command-line setting takes precedence over the configuration file setting.
The Sequence Index is a tab delimited file that starts with a header line. The columns may be in any order.
Following the header line, there is one line per single-end read and one line per paired-end read (only 1 line per pair).
Required Column Names:
- MERGE_NAME - base name for the resulting BAM file for the sample (used to group multiple fastqs or fastq pairs into a single BAM)
- FASTQ1 - name of the fastq or the first in the pair if paired-end. (Only 1 line per pair)
Optional Column Names:
- FASTQ2 - name of the 2nd fastq in paired-end reads. Specify '.' if the column exists, but this line is single-ended.
- RGID - Read Group ID for this entry
- SAMPLE - Sample Name for this entry
- LIBRARY - Library for this entry
- CENTER - Center Name for this entry
- PLATFORM - Platform for this entry
The RGID, SAMPLE, LIBRARY, CENTER, and PLATFORM are used to populate the Read Group information for this entry. These fields are optional. Either leave the column header out of the file or specify '.' if the column header exists, but the data is N/A. As long as the RGID field is specified non-N/A fields are added to the BAM file.
MERGE_NAME FASTQ1 FASTQ2 RGID SAMPLE LIBRARY CENTER PLATFORM Sample1 fastq/S1/F1_R1.fastq.gz fastq/S1/F1_R2.fastq.gz RGID1 SampleID1 Lib1 UM ILLUMINA Sample1 fastq/S1/F2_R1.fastq.gz fastq/S1/F2_R2.fastq.gz RGID1a SampleID1 Lib1 UM ILLUMINA Sample2 fastq/S2/F1_R1.fastq.gz fastq/S2/F1_R2.fastq.gz RGID2 SampleID2 Lib2 UM ILLUMINA Sample2 fastq/S2/F2.fastq.gz . RGID2 SampleID2 Lib2 UM ILLUMINA
The --fastq/FASTQ setting can be used to specify a prefix to the FASTQ1/FASTQ2 file paths that should be applied before using the files.
Reference Files
The following Reference Files are required:
- Reference File fasta files
- Files required: .fa, -bs.umfa, .GCContent, .amb, .ann, .bwt, .pac, .rbwt, .rpac, .rsa, .sa
- If you don't have the -bs.umfa file, the software will try to create it in the same directory as the reference fasta.
- .GCContent can be generated using qplot, see: QPLOT: Input Files: --gccontent and name the resulting file as
.fa.GCcontent - Use
bin/bwa index ref.faif you need to generate the bwa reference files (.amb, .ann, .bwt, .pac, .rbwt, .rpac, .rsa, .sa)
- Configuration Name: FA_REF - specify the ref.fa/ref.fa.gz name
- Files required: .fa, -bs.umfa, .GCContent, .amb, .ann, .bwt, .pac, .rbwt, .rpac, .rsa, .sa
- DBSNP File
- tab delimited file/VCF, can be compressed
- 1st column -> chromosome
- 2nd column -> 1-based position
- Configuration Name: DBSNP_VCF
- tab delimited file/VCF, can be compressed
- PLINK-compatible binary genotype files
- Files required: .bed, .bin, .fam
- Configuration Name: PLINK
Configuration File
Configuration file contains the run-time options including the software binaries and command line arguments. A default configuration file is automatically loaded. Users may specify their own configuration file specifying just the values different than the defaults. The configuration file is not required if there are no values to override.
Comments begin with a #
Format: KEY = value
Where KEY is the item being set and value is its new value
See Command-Line Options for values that can be set eitehr via command line or via configuration.
Note: Command-line options take priority over configuration file settings
Required Settings
See Reference Files for the required reference file settings.
See Sequence Index File for how to set the index file either via command line options or via configuration.
Turning Off Optional Steps
Quality Control steps can be disabled.
To Disable QPLOT, set:
RUN_QPLOT = 0
To Disable VerifyBamID, set:
RUN_VERIFY_BAM_ID = 0
Optional Configurable Settings
You may want to adjust the amount of memory/threads that are used:
There are additional configurable settings, but these are the ones most likely to be adjusted.
- BWA_THREADS = -t N
- Fill in the N with the number of threads you want BWA to run with, default is 1
- BWA_MAX_MEM = 2000000000
- Maximum amount of memory used by samtools sort after running bwa
- JAVA_MEM = -Xmx4g
- Set the maximum size of the java memory allocation pool. Default is 4g, adjust that as necessary.
Running the Mapping Pipeline
Command-Line Options
- help - print usage
- test OUTPUT_DIR - run the test example placing the output in a user specified OUTPUT_DIR. No other options are required.
- out_dir OUTPUT_DIR - directory for the output
- May also be specified via OUT_DIR in the configuration file
- Required to be set either via command-line or configuration
- conf CONFIG_FILE - configuration file
- index_file INDEX_FILE_NAME - name of the index file
- May also be specified via INDEX_FILE in the configuration file
- Required to be set either via command-line or configuration
- ref_dir REFERENCE_DIR - value to set config key REF_DIR to, overriding other values, REF_DIR can then be used inside config files.
- May also be specified via REF_DIR in the configuration file
- fastq FASTQ_PATH - prefix path to the fastq files specified in the INDEX_FILE
- May also be specified via FASTQ in the configuration file
- keepTmp - Do not remove the temporary files (removed by default)
- May also be specified via KEEP_TMP in the configuration file
- numjobs N - Replace N with the number of jobs that should be run in parallel
Note: Command-line options take priority over configuration file settings
The mapping pipeline is currently a 2 step process.
Step 1: Generate the Makefiles, 1 per sample
Run bin/gen_biopipeline.pl with the appropriate command-line parameters.
Example:
bin/gen_biopipeline.pl --conf config.txt --out_dir output
This step generates 1 Makefile per sample in the output/Makefiles/ directory, but does not run them. These Makefiles contain all of the information to run each sample.
Instructions are printed for running the Makefiles.
Step 2: Run the Samples
Run make -f on each file in the Makefiles directory. Each Makefile is independent and can be run in parallel and across a cloud.
It is recommended that you redirect stdout and stderr to files to save the results.
On failure, the Makefile should report a message like:
make: *** [...] Error 1
Where ... is filled in with other text indicating what step failed.
On success, you should see the following subdirectories under the user specified output directory:
- alignment_recal/
- Makefiles/
- QCFiles/ (if all quality control is not disabled)
- tmp/
You should see a .OK for each Sample in the index file.
If you do not see these .OK files, then your Mapping Pipeline failed.
On success, the alignment.recal/ directory contains the final BAMs, bais, and md5s.
If processing fails part way through, you can pick up where you left off by rerunning the make command.