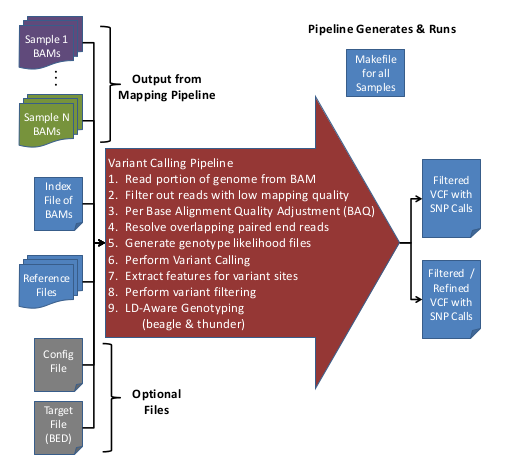
GotCloud: Variant Calling Pipeline
Back to parent: GotCloud
The Variant Calling Pipeline (UMAKE) takes recalibrated BAM files and detects SNPs and calls their genotypes, producing VCF files.
Here is an overview of the Variant Calling Pipeline:
Input Data
- Aligned/Processed/Recalibrated BAM files
- Index file containing Sample IDs & BAM file names
- Reference files
- (Optional) Configuration file to override default options
BAM files
The BAM files need to be duplicate-marked and base-quality recalibrated in order to obtain high quality SNP calls.
FASTQs can be converted to this type of BAM using the Mapping Pipeline.
Index File
Each line of the index file represents each individual under the following format. Note that multiple BAMs per individual may be provided.
[SAMPLE_ID] [COMMA SEPARATED POPULATION LABELS] [BAM_FILE1] [BAM_FILE2] ...
Columns:
- sample id
- comma separated population labels
- BAM File 1
- BAM File 2 (if applicable)
- ...
- # BAM File N
Reference Files
Reference files are required for doing Variant Calling.
- Reference Sequence in fasta format.
- Configuration File Setting:
REF = path/file.fa
- Configuration File Setting:
- Indel VCF File Prefix
- Configuration File Setting:
INDEL_PREFIX = path/indels.sites.hg19 path/containsindels.sites.hg19.chr20.vcffor each chromosome being processed
- Configuration File Setting:
- DBSNP File Prefix
- Configuration File Setting:
DBSNP_PREFIX = path/dbsnp_135_b37.rod path/containsdbsnp_135_b37.rod.chr20.mapfor each chromosome being processed
- Configuration File Setting:
- HapMap3 polymorphic site prefix
- Configuration File Setting:
HM3_PREFIX = path/hapmap3.qc.poly path/containshapmap3.qc.poly.chr20.bim&hapmap3.qc.poly.chr20.frqfor each chromosome being processed
- Configuration File Setting:
A set of reference files can be downloaded from: [| FTP Download of Full Resource Files]
Configuration File Example Reference Settings:
REF = path/file.fa INDEL_PREFIX = path/indels.sites.hg19 DBSNP_PREFIX = path/dbsnp_135_b37.rod HM3_PREFIX = path/hapmap3.qc.poly
Configuration File
Configuration file contains the run-time options including the software binaries and command line arguments. A default configuration file is automatically loaded. Users must specify their own configuration file specifying just the values different than the defaults.
Comments begin with a #
Format: KEY = value
Where KEY is the item being set and value is its new value
Required User Config Files Settings
The following Config File Settings must be specified by the user:
- CHRS = space separated list of chromosomes you want
- BAM_INDEX = path to the Index File of BAMs
Required on Command-Line or in Config File
The following Command-Line or Config File Settings must be specified by the user:
- --outdir/OUTDIR= path to desired output directory
Targeted/Exome Sequencing Settings
If you are running Targeted/Exome Sequencing, the user should specify:
- Write loci file when performing pileup
- WRITE_TARGET_LOCI = TRUE
- Specify the output sub-directory to store target information, for example: targetDir
- Should not be a full path as this will co under the OUTDIR directory.
- TARGET_DIR = targetDir
If all individuals have the same target:
- Specify the single bed file, for example: target.bed
- UNIFORM_TARGET_BED = target.bed
If not all individuals have the same target:
- Specify the file containing the sample id -> bed map, for example: targetMap.txt
- MULTIPLE_TARGET_MAP = targetMap.txt
- Each line of the file contains [SM_ID] [TARGET_BED]
- MULTIPLE_TARGET_MAP = targetMap.txt
Optional Settings:
- Extend the target region by a given number of bases, for example: 50
- OFFSET_OFF_TARGET = 50
Configure Reference Files
See Reference Files for information on how to specify the reference files.
Chromosome X Calling
- PED_INDEX = pedfile.ped
Running
Running umake is straightforward:
/usr/local/biopipe/bin/umake.pl --conf umake.conf --snpcall --numjobs 2
Replace umake.conf with the approprate path/name of the user's configuration file.
If OUTDIR is not defined in the configuration file, add --outdir followed by the path to the user's desired output directory.
Update the value following --numjobs to the appropriate number of jobs to be run in parallel.
Running on a Cluster
To run on the Cluster, the following settings need to be added to the configuration file:
SLEEP_MULT = 20 MOS_PREFIX = # PREFIX FOR MOSIX COMMAND (BLANK IF UNUSED) MOS_NODES = # COMMA-SEPARATED LIST OF NODES TO SUBMIT JOBS REMOTE_PREFIX = # REMOTE_PREFIX : Set if cluster node see the directory differently (e.g. /net/mymachine/[original-dir])
Set the MOS_NODES to the appropriate node list.
Update MOS_PREFIX to the applicable prefix.
- For MOSIX, use:
MOS_PREFIX = mosrun -E/tmp -t -i
Results
If there is a failure, you should see a message like:
make: *** [...] Error 1
Where ... is filled in with other text indicating what step failed.
On SNP Call success, you should see the following output sub-directories under your output directory:
- glfs with a bams & samples subdirectory
- pvcfs with a subdirectory per chromosome and then per region
- split with a subdirectory per chromosome
- vcfs with a subdirectory per chromosome
- (optionally your target directory)
Under the vcf/chrXX directory, there should be:
- chrXX.filtered.sites.vcf
- chrXX.filtered.sites.vcf.log
- chrXX.filtered.sites.vcf.summary
- chrXX.filtered.vcf.gz
- chrXX.filtered.vcf.gz.OK
- chrXX.filtered.vcf.gz.tbi
- chrXX.merged.sites.vcf
- chrXX.merged.stats.vcf
- chrXX.merged.vcf
- chrXX.merged.vcf.OK
The .merged.vcf is the merged together versions of the separate regions in the same chromosome.
The filtered is the merged.vcf after it has been run through filters and is marked with PASS/FAIL.
Under the split/chrXX directory, there should be:
- chrXX.filtered.PASS.split.[N].vcf.gz
- chrXX.filtered.PASS.split.err
- chrXX.filtered.PASS.split.vcflist
- chrXX.filtered.PASS.gz
- subset.OK