LASER
Introduction
LASER is a C++ software that can estimate individual ancestry directly from genome-wide shortgun sequencing reads without calling genotypes. The method relies on the availability of a set of reference individuals whose genome-wide SNP genotypes and ancestral information are known. We first construct a reference coordinate system by applying principal components analysis (PCA) to the genotype data of the reference individuals. Then, for each sequence sample, uses the genome-wide sequencing reads to place the sample into the reference PCA space. With an appropriate reference panel, the estimated coordinates of the sequence samples identify their ancestral background and can be directly used to correct for population structure in association studies or to ensure adequate matching of cases and controls.
This goal of this wiki page is to help you get start using LASESR, and we encourage you to read LASER Manual for details.
Workflow
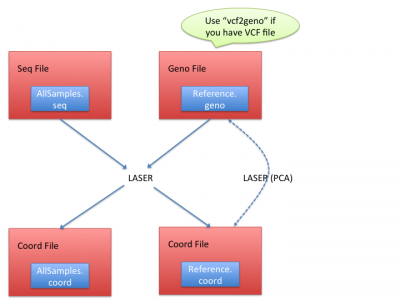
LASER generates the coordinates of reference panels and sequencing samples together, and it requires essentially two input files:
- Seq file: a text file processed from BAM (alignment) files. (See Processing sequencing file for how to obtain seq file)
- Geno file: genotypes of reference panels. (See Geno file to understand geno file format)
In coord files of reference, LASER calculated the Principal Component Analysis (PCA) results of the reference samples; In coord files of sequencing samples, LASER infers their ancestries by placing their ancestry coordinates on the reference coordinates.
An example result is shown below:
popID indivID L1 Ci t PC1 PC2 YRI NA19238 1409 0.304122 0.98933 52.7634 -39.7924 CEU NA12892 1552 0.330037 0.989709 9.82674 25.2898 CEU NA12891 1609 0.362198 0.988082 0.439573 26.8872 CEU NA12878 1579 0.334825 0.988677 8.83775 28.1342 YRI NA19239 1558 0.34898 0.988302 53.9104 -39.1727 YRI NA19240 1735 0.404142 0.990264 59.8379 -45.2765
In the header line, popID means "population ID", indivID means "individual ID", L1 means number of loci has been covered, Ci means "average coverage", t means Procrustes similarity. PC1, PC2 means coordinates of first and second principal components.
Tutorial
In this tutorial, we will show you how to prepare data and run LASER.
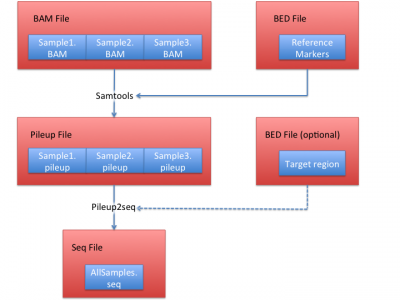
Process sequencing file (BAM)
We illustrate how to obtain .seq file from BAM files in this section. In this example, we use HGDP data set, which contain 938 individuals and 632,958 markers as the reference.
1. Obtain pileup files from BAM files
We use samtools to extract the bases on the 632,958 reference markers using:
samtools mpileup -q 30 -Q 20 -f ../../LASER-resource/reference/hs37d5.fa -l HGDP_938.bed exampleBAM/NA12878.chrom22.recal.bam > NA12878.chrom22.pileup
2. Obtain seq files from pileup files.
To convert pile up files into seq file format, we first generate site file:
cat ../resource/HGDP/HGDP_938.site |awk '{if (NR > 1) {print $1, $2-1, $2;}}' > HGDP_938.bed
Then use this site file and all generated pileup files from step 1 to generate a seq file:
python pileup2seq.py -m ../resource/HGDP/HGDP_938.site -o test NA12878.chrom22.pileup
Estimating ancestry using LASER
The easiest way to use LASER using provide example is:
./laser -s pileup2seq/test.seq -g resource/HGDP/HGDP_938.geno -c resource/HGDP/HGDP_938.RefPC.coord -o test -k 2
Upon successful running, you will find result file "test.SeqPC.coord".
File format
Geno file
In our resource folder, we provide an example geno file for the HGDP data set (resource/HGDP/HGDP_938.geno):
Brahui HGDP00001 1 2 1 1 0 2 0 2 1 2 2 2 1 1 2 1 0 Brahui HGDP00003 0 0 2 0 0 2 0 2 0 2 2 2 2 0 2 2 0 Brahui HGDP00005 0 2 2 0 0 1 0 2 1 2 2 2 2 1 2 2 1 Brahui HGDP00007 0 2 2 0 0 2 0 2 0 2 2 2 1 1 2 2 1 Brahui HGDP00009 0 1 0 1 0 2 0 2 0 2 2 2 2 0 2 2 0 Brahui HGDP00011 1 1 2 1 1 2 1 1 1 2 2 2 1 1 2 2 0 Brahui HGDP00013 1 2 2 1 1 2 1 2 0 2 2 2 2 0 2 2 0 Brahui HGDP00015 1 1 2 0 0 2 0 2 0 2 2 2 2 0 2 2 0 Brahui HGDP00017 1 1 2 0 0 1 0 0 0 2 0 1 1 2 2 2 0 Brahui HGDP00019 0 2 2 0 0 1 0 1 0 2 1 2 2 1 2 2 0
The first and second columns represent the population id and individual id. From the third column, the number represents the genotype. In this geno file, we have 632,960 columns which contains 632,956 markers from column 3 to the last column.
Seq file
Seq file organizes the sequencing information into LASER readable format. The first two columns are intended for population id and individual id. Subsequent columns are total read depth and reference base count. For example, column 3 and 4 are 0, 0 in the following example, meaning at first marker, the read depth is 0 and none of read has reference base. We enforce tab delimiters between markers and space delimiters between read depth and reference base counts. An example seq file is shown below:
NA12878.chrom22 NA12878.chrom22 0 0 0 0 0 0 0 0 0
Pileup file
Pileup file are generate by samtools. An example pileup file is listed below:
22 17094749 A 1 c D 22 17202602 T 1 . D 22 17411899 A 1 . C 22 17450515 G 2 ., 9< 22 17452966 T 1 c 5 22 17470779 C 1 , A 22 17492203 G 1 , B 22 17504945 C 3 ,.. BCA 22 17529814 T 3 .., CCC
The columns are chromosome, position (1-based), reference base, depth, bases and base qualities.
BED file
BED file represents genomic regions and it follows UCSC conventions:
1 752565 752566 1 768447 768448 1 1005805 1005806 1 1018703 1018704 1 1021414 1021415
The columns are: chromosome, start position (0-based) and end position (1-based).
Coord file
Coord files are used to represents the ancestries of both reference samples and sequencing samples. An example coord file looks like below:
popID indivID L1 Ci t PC1 PC2 YRI NA19238 1409 0.304122 0.98933 52.7634 -39.7924 CEU NA12892 1552 0.330037 0.989709 9.82674 25.2898 CEU NA12891 1609 0.362198 0.988082 0.439573 26.8872 CEU NA12878 1579 0.334825 0.988677 8.83775 28.1342 YRI NA19239 1558 0.34898 0.988302 53.9104 -39.1727 YRI NA19240 1735 0.404142 0.990264 59.8379 -45.2765
The columns are: popID means "population ID", indivID means "individual ID", L1 means number of loci has been covered, Ci means "average coverage", t means Procrustes similarity. PC1, PC2 means coordinates of first and second principal components.
Site file
Site file is equivalent to BED file and it is used here to represent marker positions. An example site file looks like below:
CHR POS ID REF ALT 1 752566 rs3094315 G A 1 768448 rs12562034 G A 1 1005806 rs3934834 C T 1 1018704 rs9442372 A G 1 1021415 rs3737728 A G
The site file has header line, and it contains chromosome, position(1-based), id (usually marker name), ref (reference allele) and alt (alternative allele).
Advanced options
LASER has advanced options including (1) running parallel jobs; (2) increase ancestry inference using repeated runs; (3) generate PCA coordiates for genotypes. See its manual for detailed information LASER Manual.
Contact
Comments on this wiki page or questions related to preparing input files for LASER can be sent to Xiaowei Zhan. \\ Comments on the LASER software or the user's manual can be sent to Chaolong Wang. \\ This project was directed by Gonçalo Abecasis and Sebastian Zöllner at the University of Michigan.