SeqShop: Calling Your Own Genome, December 2014
Login instructions for seqshop-server
Login to the seqshop-server Linux Machine
This section will appear redundantly in each session. If you are already logged in or know how to log in to the server, please skip this section
- Login to the windows machine
- The username/password for the Windows machine should be written on the right-hand monitor
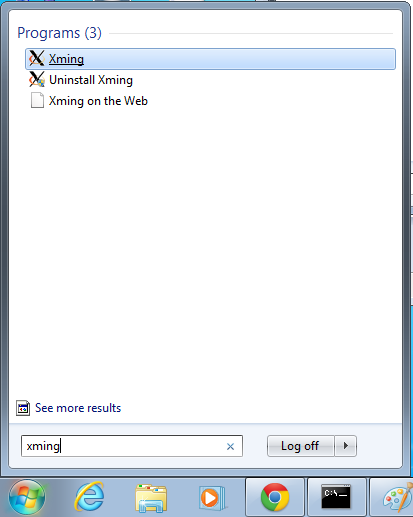
- Start xming so you can open external windows on our Linux machine
- Start->Enter "Xming" in the search and select "Xming" from the program list
- Nothing will happen, but Xming was started.
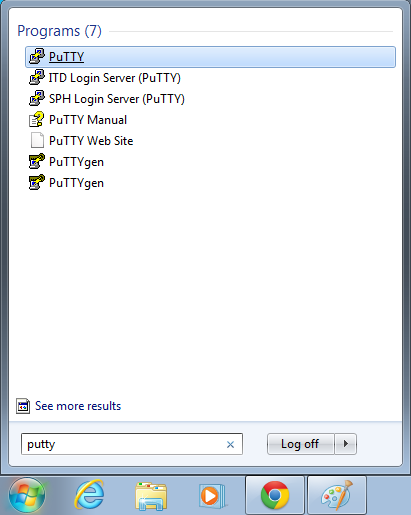
- Open putty
- Start->Enter "putty" in the search and select "PuTTY" from the program list
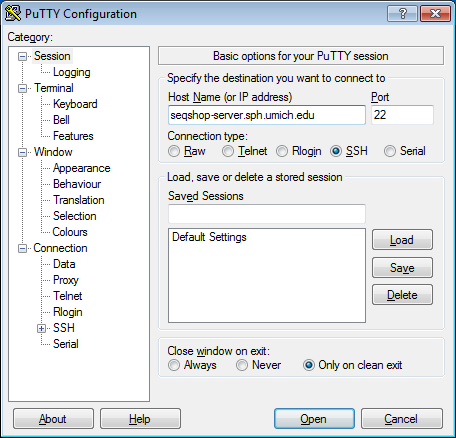
- Configure PuTTY in the PuTTY Configuration window
- Host Name:
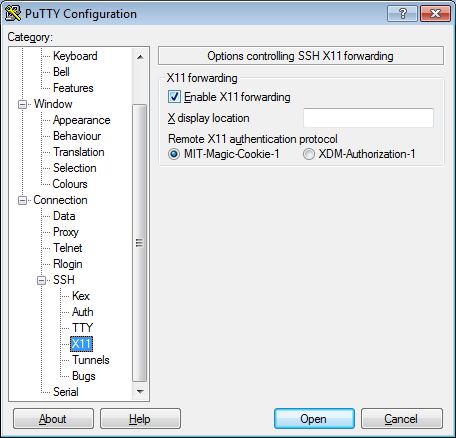
seqshop-server.sph.umich.edu - Setup to allow you to open external windows:
- In the left pannel: Connection->SSH->X11
- Add a check mark in the box next to
Enable X11 forwarding - Click
Open - If it prompts about a key, click
OK - Enter your provided username & password as provided
You should now be logged into a terminal on the seqshop-server and be able to access the test files.
- If you need another terminal, repeat from step 3.
Login to the seqshop Machine
So you can each run multiple jobs at once, we will have you run on 4 different machines within our seqshop setup.
- You can only access these machines after logging onto seqshop-server
3 users logon to:
ssh -X seqshop1
3 users logon to:
ssh -X seqshop2
2 users logon to:
ssh -X seqshop3
2 users logon to:
ssh -X seqshop4
Setup
The snpcall pipeline will run overnight, but you'll want to log out.
- How do I leave something running on the server even if I log out?
- One solution is screen!
- How do I use screen?
- Before running your command, you need to start screen:
screen
As it says, press Space or Return.
- It should now look basically the same as your normal command line.
- Scrolling problems when using screen?
- If you want to scroll and screen doesn't scroll like you normally would?
- Type Ctrl-a Esc and you should be able to scroll up with your mouse wheel
- Or at least that is what I do from my Linux machine - (sorry I'm typing this up/testing these commands from Linux and not windows, so can't test it out)
- Type Ctrl-a Esc and you should be able to scroll up with your mouse wheel
Set these values. Also, be sure to specify your sample name instead of SampleXX
export SAMPLE=SampleXX source /net/seqshop-server/home/mktrost/seqshop/setupSS.txt
See the settings you just used:
cat /net/seqshop-server/home/mktrost/seqshop/setupSS.txt
Shows you:
export GC=/net/seqshop-server/home/mktrost/seqshop/gotcloudexport OUT=~/$SAMPLE/output
SnpCall Step 1 (Day 1): Start SnpCall
List of BAMs
The list of BAMs has already been created (just 1 BAM, your sample).
- But it is simply SAMPLE\tBAM_name, so easy to figure out
cat ~/$SAMPLE/output/bam.list
SampleXX SampleXX/output/bams/SampleXX.recal.bam
- Relative path, so assumes running from your home directory (I prefer absolute paths, but for simplicity of the workshop, we just use relative path).
Configuring SnpCall
cat ~/$SAMPLE/gotcloud.conf
You will see something like this:
# Cluster Settings BATCH_TYPE = BATCH_OPTS = OUT_DIR = Sample13/output # Align Settings MAP_TYPE = BWA_MEM BWA_THREADS = -t 24 FASTQ_LIST = fastq.list # SNP Call Settings UNIT_CHUNK = 20000000 # Chunk size of SNP calling : 20Mb VCF_EXTRACT = /net/seqshop-server/home/mktrost/seqshop/singleSample/snpOnly.vcf.gz MODEL_GLFSINGLE = TRUE MODEL_SKIP_DISCOVER = FALSE MODEL_AF_PRIOR = TRUE EXT_DIR = /net/seqshop-server/home/mktrost/seqshop/singleSample/ext EXT = $(EXT_DIR)/ALL.chrCHR.phase3.combined.sites.unfiltered.vcf.gz $(EXT_DIR)/chrCHR.filtered.sites.vcf.gz
Configuration Updates
In order to complete SnpCall overnight, we are going to tell GotCloud to only call SNPs for the EXOME regions.
Edit
nedit $SAMPLE/gotcloud.conf&
- Or you can use
vioremacsor your favorite editor
Specify the target region:
UNIFORM_TARGET_BED = /net/seqshop-server/home/mktrost/seqshop/singleSample/20130108.exome.targets.bed
- See Sequencing Settings for more information on the GotCloud configuration settings for running Targeted/Exome runs.
Running SnpCall
Run GotCloud snpcall with 6 jobs running in parallel
- Why 6?
- You want to run as many as you can.
- 5 of you on the machine - 5*6 = 30 jobs will be running in parallel on that machine
${GC}/gotcloud snpcall --conf $SAMPLE/gotcloud.conf --numjobs 6
- Only need the configuration & number of threads, rest is specified within the configuration.
This will run overnight. We will check if it completed at the practical in the morning.
Log Out
- Want to log out and leave your job running?
In the screen window, type:
Ctrl-a d
(Hold down Ctrl and type 'a', let go of both and type 'd')
- This will "detach" from your screen session while your alignment continues to run.
If you have not detached from screen:
Ctrl-a d
exit PuTTY
Tuesday FEEDBACK!
Please provide feedback on the lectures/tutorials from today:
https://docs.google.com/forms/d/1n8xYxvsOq-HsabpDfGcHvwD84BYIRDx8_b-H5N3d-D8/viewform
SnpCall Step 2 (Day 2): Checking SnpCall
Logging Back in to Check Jobs
- How do you log back into screen tomorrow?
screen -r
This will resume an already running screen.
Indel Step 1 (Day 1): Start Indel Calling
List of BAMs
The list of BAMs has already been created (just 1 BAM, your sample).
- But it is simply SAMPLE\tBAM_name, so easy to figure out
cat ~/$SAMPLE/output/bam.list
SampleXX SampleXX/output/bams/SampleXX.recal.bam
- Relative path, so assumes running from your home directory (I prefer absolute paths, but for simplicity of the workshop, we just use relative path).
Configuring Indel
No special Configuration settings for Indel calling
cat ~/$SAMPLE/gotcloud.conf
You will see something like this:
# Cluster Settings BATCH_TYPE = BATCH_OPTS = OUT_DIR = Sample13/output # Align Settings MAP_TYPE = BWA_MEM BWA_THREADS = -t 24 FASTQ_LIST = fastq.list # SNP Call Settings UNIT_CHUNK = 20000000 # Chunk size of SNP calling : 20Mb VCF_EXTRACT = /net/seqshop-server/home/mktrost/seqshop/singleSample/snpOnly.vcf.gz MODEL_GLFSINGLE = TRUE MODEL_SKIP_DISCOVER = FALSE MODEL_AF_PRIOR = TRUE EXT_DIR = /net/seqshop-server/home/mktrost/seqshop/singleSample/ext EXT = $(EXT_DIR)/ALL.chrCHR.phase3.combined.sites.unfiltered.vcf.gz $(EXT_DIR)/chrCHR.filtered.sites.vcf.gz
Running Indel Calling
Run GotCloud indel with 6 jobs running in parallel
- Why 6?
- You want to run as many as you can.
- 5 of you on the machine - 5*6 = 30 jobs will be running in parallel on that machine
${GC}/gotcloud indel --conf $SAMPLE/gotcloud.conf --numjobs 6
- Only need the configuration & number of threads, rest is specified within the configuration.
This will run overnight. We will check if it completed at the practical in the morning.
Log Out
- Want to log out and leave your job running?
In the screen window, type:
Ctrl-a d
(Hold down Ctrl and type 'a', let go of both and type 'd')
- This will "detach" from your screen session while your alignment continues to run.
If you have not detached from screen:
Ctrl-a d
exit PuTTY
Tuesday FEEDBACK!
Please provide feedback on the lectures/tutorials from today:
https://docs.google.com/forms/d/1n8xYxvsOq-HsabpDfGcHvwD84BYIRDx8_b-H5N3d-D8/viewform
Indel Step 2 (Day 2): Checking Indel
Logging Back in to Check Jobs
- How do you log back into screen tomorrow?
screen -r
This will resume an already running screen.
Looking at INDEL results
Look in the output directory
ls ~/$SAMPLE/output
What in that directory was produced by indel calling?
gotcloud.indel.conf- dump of all configuration settings for this run
gotcloud.indel.Makefile- Makefile that was generated to manage all of the commands to be run
gotcloud.indel.Makefile.log- log of all commands run by the Makefile
indel/- indel output directory
Let's look at the indel output
ls ~/$SAMPLE/output/indel
- 3 directories
- aux - intermediate files
- indelvcf - intermediate files
- final indel files
Final indel directory:
ls ~/$SAMPLE/output/indel/final
- all.genotypes.vcf.gz - output VCF
- all.genotypes.vcf.gz.tbi - output VCF index file to allow jumping to positions
- all.genotypes.vcf.gz.tbi.OK - completion indicator
- merge/ - directory with per chromosome bcf (binary vcf) files
- all.genotypes.vcf.gz.OK - completion indicator
- all.genotypes.vcf.gz.tbi.log - log
- concat.log - log
Looking at final INDEL VCF
Check the number of passing INDEL variants:
$GC/bin/vt peek ~/$SAMPLE/output/indel/final/all.genotypes.vcf.gz -f "PASS"
Gives something like:
no. Indels : 570661
2 alleles (ins/del) : 570661 (0.87) [265448/305213]
>=3 alleles (ins/del) : 0 (-nan) [0/0]
Check the number of passing INDEL's with allele count > 0:
$GC/bin/vt peek ~/$SAMPLE/output/indel/final/all.genotypes.vcf.gz -f "PASS&&INFO.AC!=0"
Gives something like:
no. Indels : 549963
2 alleles (ins/del) : 549963 (0.91) [261480/288483]
>=3 alleles (ins/del) : 0 (-nan) [0/0]
Some INDELs had allele count 0.
Check the number of passing INDEL's with allele count 2:
$GC/bin/vt peek ~/$SAMPLE/output/indel/final/all.genotypes.vcf.gz -f "PASS&&INFO.AC==2"
Gives something like:
no. Indels : 216134
2 alleles (ins/del) : 216134 (1.17) [116511/99623]
>=3 alleles (ins/del) : 0 (-nan) [0/0]
Check the number of passing INDEL's with allele balance > 0.5:
$GC/bin/vt peek ~/$SAMPLE/output/indel/final/all.genotypes.vcf.gz -f "PASS&&INFO.AB>0.5"
Gives something like:
no. Indels : 132878
2 alleles (ins/del) : 132878 (0.68) [53714/79164]
>=3 alleles (ins/del) : 0 (-nan) [0/0]
Check the number of passing INDEL's with allele balance < 0.5:
$GC/bin/vt peek ~/$SAMPLE/output/indel/final/all.genotypes.vcf.gz -f "PASS&&INFO.AB<0.5"
Gives something like:
no. Indels : 169198
2 alleles (ins/del) : 169198 (0.89) [79504/89694]
>=3 alleles (ins/del) : 0 (-nan) [0/0]