SeqShop: Estimates of Genetic Ancestry Practical, June 2014
Introduction
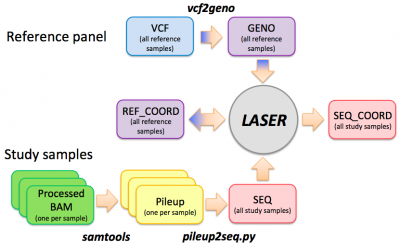
See the tutorial slides for an introduction of the LASER analysis workflow, input/output file formats, and usage of the LASER software.
The main purpose of this page is to provide step-by-step command lines for estimating ancestry of 6 targeted sequencing samples (2 HapMap trios) in a principal component space generated using genome-wide SNP data from the Human Genome Diversity Project (HGDP).
For reference of options and usage of the software, please read the manual.
LASER workflow
Getting started
Create a working directory:
mkdir ancestry cd ancestry
Download and decompress software package:
wget http://www.sph.umich.edu/csg/chaolong/LASER/LASER-2.01.tar.gz tar xzvf LASER-2.01.tar.gz
Set up to access data:
source /home/chaolong/LASER-Tutorial/setup.txt
What is in the setup.txt file:
export GC=/home/mktrost/seqshop/gotcloud export BAM=/home/chaolong/LASER-Tutorial/BAM export REF=/home/chaolong/LASER-Tutorial/reference export HGDP=/home/chaolong/LASER-Tutorial/HGDP
Preparing input files for LASER
Step 0: vcf --> geno
This step prepares the reference panel by converting a VCF genotype file to a GENO file. We will skip this step and use a ready-to-use HGDP reference panel. A typical command to run the vcf2geno tool is given in the file "./LASER-2.01/vcf2geno/cmd.sh".
Step 1: bam --> pileup
This step uses samtools to generate pileup files from bam files. Please only try one sample so that we won't overload the sever with everyone running 6 jobs at the same time. Pileup files for these 6 samples have been prepared for later steps. It takes about 2 mins for each pileup job.
$GC/bin/samtools mpileup -q 30 -Q 20 -f ./reference/hs37d5.fa.rz -l ./HGDP/HGDP_938.bed ./BAM/121101035.recal.bam > 121101035.recal.pileup & # $GC/bin/samtools mpileup -q 30 -Q 20 -f ./reference/hs37d5.fa.rz -l ./HGDP/HGDP_938.bed ./BAM/121101043.recal.bam > 121101043.recal.pileup & # $GC/bin/samtools mpileup -q 30 -Q 20 -f ./reference/hs37d5.fa.rz -l ./HGDP/HGDP_938.bed ./BAM/121101050.recal.bam > 121101050.recal.pileup & # $GC/bin/samtools mpileup -q 30 -Q 20 -f ./reference/hs37d5.fa.rz -l ./HGDP/HGDP_938.bed ./BAM/121101052.recal.bam > 121101052.recal.pileup & # $GC/bin/samtools mpileup -q 30 -Q 20 -f ./reference/hs37d5.fa.rz -l ./HGDP/HGDP_938.bed ./BAM/121101415.recal.bam > 121101415.recal.pileup & # $GC/bin/samtools mpileup -q 30 -Q 20 -f ./reference/hs37d5.fa.rz -l ./HGDP/HGDP_938.bed ./BAM/121101861.recal.bam > 121101861.recal.pileup &
Step 2: pileup --> seq
This step will generate a file called "hapmap_trios.seq", containing the information of 6 samples. It takes about 30 seconds to run. We used the pre-generated pileup files in the $BAM folder.
python ./LASER-2.01/pileup2seq/pileup2seq.py \ -m $HGDP/HGDP_938.site \ -b $BAM/AMD_roi_1-based.bed \ -i $BAM/AMD_hapmap_trios_id.txt \ -o hapmap_trios \ $BAM/121101035.recal.pileup \ $BAM/121101043.recal.pileup \ $BAM/121101050.recal.pileup \ $BAM/121101052.recal.pileup \ $BAM/121101415.recal.pileup \ $BAM/121101861.recal.pileup &
In the above command, -i specifies alternative IDs for the BAM files to be used in the .seq file (including popID and indivID). -b and -i are optional.