Difference between revisions of "SeqShop: Calling Your Own Genome, May 2015"
| (33 intermediate revisions by 2 users not shown) | |||
| Line 102: | Line 102: | ||
</div> | </div> | ||
| − | <div class="mw-collapsible" style="width:500px"> | + | <div class="mw-collapsible mw-collapsed" style="width:500px"> |
== Thursday == | == Thursday == | ||
| Line 156: | Line 156: | ||
=== FEEDBACK!=== | === FEEDBACK!=== | ||
Please provide feedback for today. | Please provide feedback for today. | ||
| − | https://docs.google.com/forms/d/ | + | https://docs.google.com/a/umich.edu/forms/d/1iES6usHxLB7Ec9hRxtqYgH7v05lU3Ume4VJcksx8Ogg/viewform |
</div> | </div> | ||
| Line 163: | Line 163: | ||
</div> | </div> | ||
| − | <div class="mw-collapsible | + | <div class="mw-collapsible" style="width:1000px"> |
== Friday == | == Friday == | ||
| − | <div class="mw-collapsible-content | + | <div class="mw-collapsible-content"> |
| Line 172: | Line 172: | ||
[[SeqShop: Ancestry On Your Own Genome, May 2015]] | [[SeqShop: Ancestry On Your Own Genome, May 2015]] | ||
| − | === | + | === Setup Variables === |
| − | + | Set these values. Also, be sure to specify your sample name instead of SampleXX | |
| + | export SAMPLE=SampleXX | ||
| + | or | ||
| + | export SAMPLE=NA12878 | ||
| − | + | Point to your GotCloud & your output directory: | |
| − | + | export GC=~/seqshop/gotcloud | |
| − | + | export OUT=~/$SAMPLE/output | |
| − | |||
| − | |||
| − | |||
=== Reviewing Indel Results === | === Reviewing Indel Results === | ||
| − | |||
| − | |||
| − | |||
| − | |||
Look in the output directory | Look in the output directory | ||
ls ~/$SAMPLE/output | ls ~/$SAMPLE/output | ||
| Line 251: | Line 247: | ||
The insertion deletion ratio increases from 0.91 to 0.92. | The insertion deletion ratio increases from 0.91 to 0.92. | ||
| + | === Return to SeqShop: Ancestry On Your Own Genome, May 2015 === | ||
| + | Return to [[SeqShop:_Ancestry_On_Your_Own_Genome,_May_2015#Checking_if_Pileup_finished]] | ||
=== Friday: Reviewing SNPCALL Results === | === Friday: Reviewing SNPCALL Results === | ||
| Line 260: | Line 258: | ||
=== Friday : More SNP Analysis === | === Friday : More SNP Analysis === | ||
| + | In addition, set another environmental variable for locating the binaries for custom analysis | ||
| − | ==== | + | export HK=/net/seqshop-server/home/hmkang/apigenome/bin |
| + | export EPACTS=/net/seqshop-server/home/mktrost/seqshop/epacts/ | ||
| + | export REF=/net/seqshop-server/home/mktrost/seqshop/singleSample/ref/gotcloud.ref | ||
| + | export GC=~/seqshop/gotcloud | ||
| − | |||
| − | + | export SAMPLE=SampleXX | |
| − | export SAMPLE=SampleXX | + | export OUT=~/$SAMPLE/output |
| − | |||
| − | |||
| − | |||
| − | |||
| − | export | ||
==== Annotation / Lookup against dbSNP ==== | ==== Annotation / Lookup against dbSNP ==== | ||
| Line 277: | Line 273: | ||
If you want to add rsIDs to your variant files, you can do this by running the following command | If you want to add rsIDs to your variant files, you can do this by running the following command | ||
| − | $HK/vcf-add-rsid -vcf $OUT/vcfs/chr1/chr1.filtered.vcf.gz --db $HK/../data/ | + | $HK/vcf-add-rsid -vcf $OUT/vcfs/chr1/chr1.filtered.vcf.gz --db $HK/../data/dbsnp_142.b37.vcf.gz --out $OUT/vcfs/chr1/chr1.filtered.rsid.vcf.gz |
If you want to run this command across all chromosomes in parallel, you can use the special script run-command-wgs | If you want to run this command across all chromosomes in parallel, you can use the special script run-command-wgs | ||
| − | $HK/run- | + | $HK/run-make --repeat-chr --cmd "$HK/vcf-add-rsid -vcf $OUT/vcfs/chr1/chr1.filtered.vcf.gz --db $HK/../data/dbsnp_142.b37.vcf.gz --out $OUT/vcfs/chr1/chr1.filtered.rsid.vcf.gz" --numjobs 6 --out runmake.rsid |
| − | Looking up SNPs by rsID is possible by (for example) | + | Looking up SNPs by rsID is possible by (for example, rs17766217) -- How can we find its position? |
| − | $HK/ | + | $HK/tabix $OUT/vcfs/chr8/chr8.filtered.rsid.vcf.gz 8:128504497 | less |
* Be sure to look at the QUAL & your sample's PL, and not just the GL field. Check if QUAL is 0 or PL is 0,0,0 - NS is also probably 0; DP is probably 0. That means you probably didn't have any copies, so your GT may not be correct/is unknown. | * Be sure to look at the QUAL & your sample's PL, and not just the GL field. Check if QUAL is 0 or PL is 0,0,0 - NS is also probably 0; DP is probably 0. That means you probably didn't have any copies, so your GT may not be correct/is unknown. | ||
If you want to browse the rsIDs of known GWAS SNPs, you can do this by | If you want to browse the rsIDs of known GWAS SNPs, you can do this by | ||
| − | cut -f 1,8,22 $HK/../data/gwascatalog/gwascatalog.txt | | + | cut -f 1,8,12,13,22 $HK/../data/gwascatalog/gwascatalog.txt | grep -w rs17766217 |
==== Annotating your genome ==== | ==== Annotating your genome ==== | ||
| Line 296: | Line 292: | ||
Or you can run multiple chromosomes in parallel in one command | Or you can run multiple chromosomes in parallel in one command | ||
| − | $HK/run- | + | $HK/run-make --repeat-chr --cmd "$EPACTS/bin/epacts anno --in $OUT/vcfs/chr1/chr1.filtered.rsid.vcf.gz --out $OUT/vcfs/chr1/chr1.filtered.rsid.anno.vcf.gz" --numjobs 6 --out runmake.anno |
| − | + | ||
==== Extracting only exonic SNPs ==== | ==== Extracting only exonic SNPs ==== | ||
If you want to look at the exonic SNPs, you can extract using the following command | If you want to look at the exonic SNPs, you can extract using the following command | ||
| − | $HK/run- | + | $HK/run-make --repeat-chr --cmd "($HK/tabix -H $OUT/vcfs/chr1/chr1.filtered.rsid.anno.vcf.gz; zcat $OUT/vcfs/chr1/chr1.filtered.rsid.anno.vcf.gz | grep Exon;)| $HK/bgzip -c > $OUT/vcfs/chr1/chr1.filtered.rsid.anno.exon.vcf.gz" --numjobs 6 --out runmake.exome |
And they can be combined as follows | And they can be combined as follows | ||
(zcat $OUT/vcfs/chr1/chr1.filtered.rsid.anno.exon.vcf.gz; zcat $OUT/vcfs/chr[2-9]/chr*.filtered.rsid.anno.exon.vcf.gz $OUT/vcfs/chr??/chr*.filtered.rsid.anno.exon.vcf.gz $OUT/vcfs/chrX/chrX.filtered.rsid.anno.exon.vcf.gz | grep -v ^#) | $HK/bgzip -c > $OUT/wgs.filtered.rsid.anno.exon.vcf.gz | (zcat $OUT/vcfs/chr1/chr1.filtered.rsid.anno.exon.vcf.gz; zcat $OUT/vcfs/chr[2-9]/chr*.filtered.rsid.anno.exon.vcf.gz $OUT/vcfs/chr??/chr*.filtered.rsid.anno.exon.vcf.gz $OUT/vcfs/chrX/chrX.filtered.rsid.anno.exon.vcf.gz | grep -v ^#) | $HK/bgzip -c > $OUT/wgs.filtered.rsid.anno.exon.vcf.gz | ||
| + | $HK/tabix -pvcf $OUT/wgs.filtered.rsid.anno.exon.vcf.gz | ||
==== Exonic Variants NOT found by 1000G ==== | ==== Exonic Variants NOT found by 1000G ==== | ||
| Line 328: | Line 325: | ||
Want to see this from the BAM file? Use samtools tview: | Want to see this from the BAM file? Use samtools tview: | ||
| − | $GC/bin/samtools tview $SAMPLE/output/bams/$SAMPLE.recal.bam $ | + | $GC/bin/samtools tview $SAMPLE/output/bams/$SAMPLE.recal.bam $REF/hs37d5.fa |
Use 'g' & enter the Chr:Pos | Use 'g' & enter the Chr:Pos | ||
* Some patterns may indicate not real variants. | * Some patterns may indicate not real variants. | ||
| Line 337: | Line 334: | ||
The phred score at the last column quantifies the degree of functional significance | The phred score at the last column quantifies the degree of functional significance | ||
| − | |||
</div> | </div> | ||
</div> | </div> | ||
| + | |||
| + | == OVERALL COURSE FEEDBACK! == | ||
| + | Please provide feedback: | ||
| + | https://docs.google.com/forms/d/1pxfPXKwWfA71ZJM99Sevs3MwAUz2UbHAR8dnRI-kRNM/viewform | ||
Latest revision as of 12:10, 22 May 2015
Login instructions for seqshop-server
Login to the seqshop-server Linux Machine
This section will appear redundantly in each session. If you are already logged in or know how to log in to the server, please skip this section
- Login to the windows machine
- The username/password for the Windows machine should be written on the right-hand monitor
- Start xming so you can open external windows on our Linux machine
- Start->Enter "Xming" in the search and select "Xming" from the program list
- Nothing will happen, but Xming was started.
- Open putty
- Start->Enter "putty" in the search and select "PuTTY" from the program list
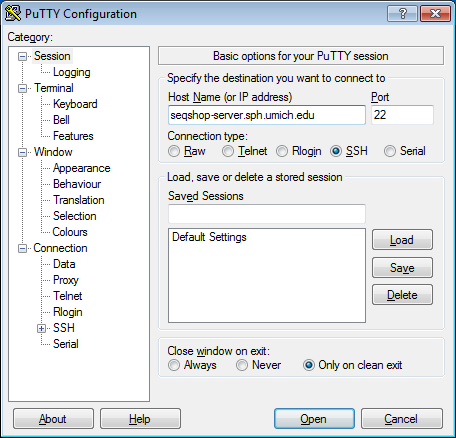
- Configure PuTTY in the PuTTY Configuration window
- Host Name:
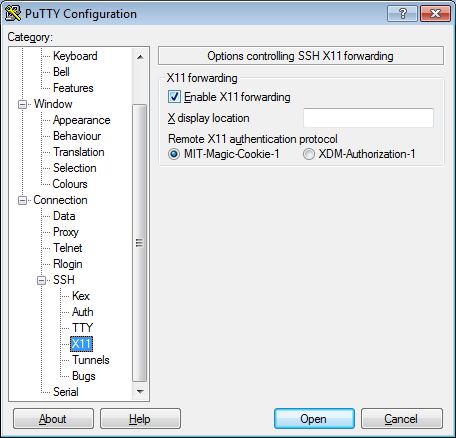
seqshop-server.sph.umich.edu - Setup to allow you to open external windows:
- In the left pannel: Connection->SSH->X11
- Add a check mark in the box next to
Enable X11 forwarding - Click
Open - If it prompts about a key, click
OK - Enter your provided username & password as provided
You should now be logged into a terminal on the seqshop-server and be able to access the test files.
- If you need another terminal, repeat from step 3.
Login to the seqshop Machine
So you can each run multiple jobs at once, we will have you run on 4 different machines within our seqshop setup.
- You can only access these machines after logging onto seqshop-server
3 users logon to:
ssh -X seqshop1
3 users logon to:
ssh -X seqshop2
2 users logon to:
ssh -X seqshop3
2 users logon to:
ssh -X seqshop4
Tuesday - Start SNP Calling
Setup Screen
The snpcall pipeline will run overnight, but you'll want to log out.
- How do I leave something running on the server even if I log out?
- One solution is screen!
- How do I use screen?
- Before running your command, you need to start screen:
screen
As it says, press Space or Return.
- It should now look basically the same as your normal command line.
Setup Variables
Set these values. Also, be sure to specify your sample name instead of SampleXX
export SAMPLE=SampleXX
or
export SAMPLE=NA12878
Point to your GotCloud & your output directory:
export GC=~/seqshop/gotcloud export OUT=~/$SAMPLE/output
List of BAMs
The list of BAMs has already been created (just 1 BAM, your sample).
- But it is simply SAMPLE\tBAM_name, so easy to figure out
cat ~/$SAMPLE/output/bam.list
SampleXX SampleXX/output/bams/SampleXX.recal.bam
- Relative path, so assumes running from your home directory (I prefer absolute paths, but for simplicity of the workshop, we just use relative path).
Configuring SNPCALL
cat ~/$SAMPLE/gotcloud.conf
You will see this:
# References SS_DIR = /net/seqshop-server/home/mktrost/seqshop/singleSample REF_DIR = $(SS_DIR)/ref/gotcloud.ref/ ######### ALIGNMENT ######## MAP_TYPE = BWA_MEM FASTQ_LIST = fastq.list BATCH_TYPE = BATCH_OPTS = BWA_THREADS = -t 6 # SNP Call Settings UNIT_CHUNK = 20000000 # Chunk size of SNP calling : 20Mb VCF_EXTRACT = $(SS_DIR)/snpOnly.vcf.gz MODEL_GLFSINGLE = TRUE MODEL_SKIP_DISCOVER = FALSE MODEL_AF_PRIOR = TRUE EXT_DIR = $(SS_DIR)/ext EXT = $(EXT_DIR)/ALL.chrCHR.phase3.combined.sites.unfiltered.vcf.gz $(EXT_DIR)/chrCHR.filtered.sites.vcf.gz
Running SNP Calling
Run GotCloud snpcall with 8 jobs running in parallel
- Why 8?
- You want to run as many as you can.
- 2-3 of you on the machine - 3*8 = 24 jobs will be running in parallel on that machine
${GC}/gotcloud snpcall --conf $SAMPLE/gotcloud.conf --numjobs 8 --outdir $OUT
- Only need the configuration, number of threads, and the output directory, rest is specified within the configuration.
This will run overnight. We will check if it completed at the practical in the morning.
Log Out
- Want to log out and leave your job running?
In the screen window, type:
Ctrl-a d
(Hold down Ctrl and type 'a', let go of both and type 'd')
- This will "detach" from your screen session while your alignment continues to run.
If you have not detached from screen:
Ctrl-a d
exit PuTTY
FEEDBACK!
Please provide feedback on today:
https://docs.google.com/forms/d/1ADTkBjzT-QNj2lrejyqGqDaahTponrw20kSgDNwqwH4/viewform
Thursday
Checking if snpcall Completed
Resume screen to Check Jobs
- How do you log back into screen?
screen -r
This will resume an already running screen.
Your screen session still has your environment variables set, so you do not need to reset them.
Verify you got a "completed successfully" message.
How long did snpcall calling take? Look at the log message - time in seconds.
List of BAMs
The list of BAMs has already been created (just 1 BAM, your sample).
- But it is simply SAMPLE\tBAM_name, so easy to figure out
cat ~/$SAMPLE/output/bam.list
SampleXX SampleXX/output/bams/SampleXX.recal.bam
- Relative path, so assumes running from your home directory (I prefer absolute paths, but for simplicity of the workshop, we just use relative path).
GotCloud INDEL Configuration
cat ~/$SAMPLE/gotcloud.conf
Same as it looked the other day with no special Configuration settings for INDEL calling.
Running INDEL
Run GotCloud indel with 6 jobs running in parallel
${GC}/gotcloud indel --conf $SAMPLE/gotcloud.conf --numjobs 6 --outdir $OUT
- Only need the configuration, number of threads, and the output directory, rest is specified within the configuration.
This will run overnight. We will check if it completed at the practical in the morning.
Log Out
- Want to log out and leave your job running?
In the screen window, type:
Ctrl-a d
(Hold down Ctrl and type 'a', let go of both and type 'd')
- This will "detach" from your screen session while your alignment continues to run.
exit PuTTY
FEEDBACK!
Please provide feedback for today. https://docs.google.com/a/umich.edu/forms/d/1iES6usHxLB7Ec9hRxtqYgH7v05lU3Ume4VJcksx8Ogg/viewform
Friday
SeqShop: Ancestry On Your Own Genome, May 2015
SeqShop: Ancestry On Your Own Genome, May 2015
Setup Variables
Set these values. Also, be sure to specify your sample name instead of SampleXX
export SAMPLE=SampleXX
or
export SAMPLE=NA12878
Point to your GotCloud & your output directory:
export GC=~/seqshop/gotcloud export OUT=~/$SAMPLE/output
Reviewing Indel Results
Look in the output directory
ls ~/$SAMPLE/output
What in that directory was produced by indel calling?
gotcloud.indel.conf- dump of all configuration settings for this run
gotcloud.indel.Makefile- Makefile that was generated to manage all of the commands to be run
gotcloud.indel.Makefile.log- log of all commands run by the Makefile
indel/- indel output directory
Let's look at the indel output
ls ~/$SAMPLE/output/indel
- 3 directories
- aux - intermediate files
- indelvcf - intermediate files
- final indel files
Final indel directory:
ls ~/$SAMPLE/output/indel/final
- all.genotypes.vcf.gz - output VCF
- all.genotypes.vcf.gz.tbi - output VCF index file to allow jumping to positions
- all.genotypes.vcf.gz.tbi.OK - completion indicator
- merge/ - directory with per chromosome bcf (binary vcf) files
- all.genotypes.vcf.gz.OK - completion indicator
- all.genotypes.vcf.gz.tbi.log - log
- concat.log - log
Looking at final INDEL VCF
Note that because this is a single sample calling, many of the INFO fields are less meaningful as many of the values like HWE p values, allele frequencies, inbreeding coefficient are a function of a population. Nonetheless, we may examine the results. First, we see how many indels were discovered for your genome:
$GC/bin/vt peek ~/$SAMPLE/output/indel/final/all.genotypes.vcf.gz
no. Indels : 588566
2 alleles (ins/del) : 588566 (0.87) [273261/315305]
This gives use 588,566 indels with an insertion deletion ratio of 0.87.
We next look at the filtered set. The PASS filter reduces the setof indels to a non overlapping set and the INFO.AC!=0 extracts all indels that are either heterozygous or homozygous alternative. Some indels that were originally discovered were found to be the homozygous reference genotype. Invariably, these are relative high depth calls where the alternative allele is discovered less or is mis-specified.
$GC/bin/vt peek ~/$SAMPLE/output/indel/final/all.genotypes.vcf.gz -f "PASS&&INFO.AC!=0"
no. Indels : 549963
2 alleles (ins/del) : 549963 (0.91) [261480/288483]
About 38K indels were removed, the insertion deletion ratio increases to 0.91. Note that in general, for high depth data, discovered indels are reported with insertion deletion ratios close to 1. So this is a good sign. Next generation sequencing errors are bias for deletions.
It is possible to perform a slightly more stringent filtering using allele balance. The allele balance estimator in this case is meaningful still for an individual because it is a function of read depth. Note that AB>0.5 denotes reference bias and AB<0.5 denotes alternative allele bias.
$GC/bin/vt peek ~/$SAMPLE/output/indel/final/all.genotypes.vcf.gz -f "PASS&&INFO.AC>0&&INFO.AB<0.7&&INFO.AB>0.3"
no. Indels : 490965
2 alleles (ins/del) : 490965 (0.92) [235254/255711]
The insertion deletion ratio increases from 0.91 to 0.92.
Return to SeqShop: Ancestry On Your Own Genome, May 2015
Return to SeqShop:_Ancestry_On_Your_Own_Genome,_May_2015#Checking_if_Pileup_finished
Friday: Reviewing SNPCALL Results
Look in the output directory
ls ~/$SAMPLE/output
Look at the vcfs:
ls ~/$SAMPLE/output/vcfs
Friday : More SNP Analysis
In addition, set another environmental variable for locating the binaries for custom analysis
export HK=/net/seqshop-server/home/hmkang/apigenome/bin export EPACTS=/net/seqshop-server/home/mktrost/seqshop/epacts/ export REF=/net/seqshop-server/home/mktrost/seqshop/singleSample/ref/gotcloud.ref export GC=~/seqshop/gotcloud
export SAMPLE=SampleXX export OUT=~/$SAMPLE/output
Annotation / Lookup against dbSNP
If you want to add rsIDs to your variant files, you can do this by running the following command
$HK/vcf-add-rsid -vcf $OUT/vcfs/chr1/chr1.filtered.vcf.gz --db $HK/../data/dbsnp_142.b37.vcf.gz --out $OUT/vcfs/chr1/chr1.filtered.rsid.vcf.gz
If you want to run this command across all chromosomes in parallel, you can use the special script run-command-wgs
$HK/run-make --repeat-chr --cmd "$HK/vcf-add-rsid -vcf $OUT/vcfs/chr1/chr1.filtered.vcf.gz --db $HK/../data/dbsnp_142.b37.vcf.gz --out $OUT/vcfs/chr1/chr1.filtered.rsid.vcf.gz" --numjobs 6 --out runmake.rsid
Looking up SNPs by rsID is possible by (for example, rs17766217) -- How can we find its position?
$HK/tabix $OUT/vcfs/chr8/chr8.filtered.rsid.vcf.gz 8:128504497 | less
- Be sure to look at the QUAL & your sample's PL, and not just the GL field. Check if QUAL is 0 or PL is 0,0,0 - NS is also probably 0; DP is probably 0. That means you probably didn't have any copies, so your GT may not be correct/is unknown.
If you want to browse the rsIDs of known GWAS SNPs, you can do this by
cut -f 1,8,12,13,22 $HK/../data/gwascatalog/gwascatalog.txt | grep -w rs17766217
Annotating your genome
You can annotate your genome using EPACTS software packages. Individual chromosome can be annotated by running.
$EPACTS/bin/epacts anno --in $OUT/vcfs/chr1/chr1.filtered.rsid.vcf.gz --out $OUT/vcfs/chr1/chr1.filtered.rsid.anno.vcf.gz
Or you can run multiple chromosomes in parallel in one command
$HK/run-make --repeat-chr --cmd "$EPACTS/bin/epacts anno --in $OUT/vcfs/chr1/chr1.filtered.rsid.vcf.gz --out $OUT/vcfs/chr1/chr1.filtered.rsid.anno.vcf.gz" --numjobs 6 --out runmake.anno
Extracting only exonic SNPs
If you want to look at the exonic SNPs, you can extract using the following command
$HK/run-make --repeat-chr --cmd "($HK/tabix -H $OUT/vcfs/chr1/chr1.filtered.rsid.anno.vcf.gz; zcat $OUT/vcfs/chr1/chr1.filtered.rsid.anno.vcf.gz | grep Exon;)| $HK/bgzip -c > $OUT/vcfs/chr1/chr1.filtered.rsid.anno.exon.vcf.gz" --numjobs 6 --out runmake.exome
And they can be combined as follows
(zcat $OUT/vcfs/chr1/chr1.filtered.rsid.anno.exon.vcf.gz; zcat $OUT/vcfs/chr[2-9]/chr*.filtered.rsid.anno.exon.vcf.gz $OUT/vcfs/chr??/chr*.filtered.rsid.anno.exon.vcf.gz $OUT/vcfs/chrX/chrX.filtered.rsid.anno.exon.vcf.gz | grep -v ^#) | $HK/bgzip -c > $OUT/wgs.filtered.rsid.anno.exon.vcf.gz $HK/tabix -pvcf $OUT/wgs.filtered.rsid.anno.exon.vcf.gz
Exonic Variants NOT found by 1000G
If you are interested in rare variants that are not identified by 1000G, you can extract them by running
zcat $OUT/wgs.filtered.rsid.anno.exon.vcf.gz | grep "EXTFILTER=NA,NA" | grep -v -w "0/0" | less
For example,
zcat $OUT/wgs.filtered.rsid.anno.exon.vcf.gz | grep "EXTFILTER=NA,NA" | grep -v -w "0/0" | perl -lane 'print "$1\t$F[6]" if ( /ANNO=([^;:]+)/)' | sort | uniq -c
will give you the counts of variants, separate by the filtering results
- Q1. How manny novel silent, missense, and nonsense SNPs are found? Is that too few, too small, or just about right?
- Q2. Looking at each functional category, which functional categories has largest fraction of SNPs failed filter? Why do you think it is?
- Q3. Can you exclude the sites that are also in dbSNP, and count how many nonsense variants are left?
To also exclude those in dbsnp:
zcat $OUT/wgs.filtered.rsid.anno.exon.vcf.gz | grep "EXTFILTER=NA,NA" | grep -v -w "0/0" | grep -v rs| perl -lane 'print "$1\t$F[6]" if ( /ANNO=([^;:]+)/)' | sort | uniq -c
Exclude dbsnp and look at Stop_Gain variants
zcat $OUT/wgs.filtered.rsid.anno.exon.vcf.gz | grep "EXTFILTER=NA,NA" | grep -v -w "0/0" |grep -v rs | perl -lane 'print "$_" if ( /ANNO=Stop_Gain/)' |grep -w PASS
Want to see this from the BAM file? Use samtools tview:
$GC/bin/samtools tview $SAMPLE/output/bams/$SAMPLE.recal.bam $REF/hs37d5.fa
Use 'g' & enter the Chr:Pos
- Some patterns may indicate not real variants.
If you want to know predicted functional significance of a particular variant, you can search by
$HK/tabix $HK/../data/CADD/whole_genome_SNVs.tsv.gz [chr]:[pos] | head -3
The phred score at the last column quantifies the degree of functional significance
OVERALL COURSE FEEDBACK!
Please provide feedback: https://docs.google.com/forms/d/1pxfPXKwWfA71ZJM99Sevs3MwAUz2UbHAR8dnRI-kRNM/viewform