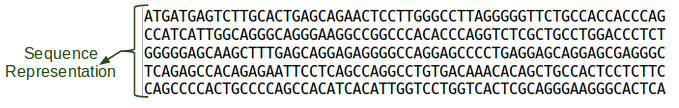Difference between revisions of "SeqShop: Sequence Mapping and Assembly Practical, June 2014"
| Line 28: | Line 28: | ||
Look at FASTQ: | Look at FASTQ: | ||
| − | less -S ${GC}/inputs/fastq/HG00641.chr7.CFTR. | + | less -S ${GC}/inputs/fastq/HG00641.chr7.CFTR.SRR069531_1.fastq |
<code>less</code> is a Linux command that allows you to look at a file. | <code>less</code> is a Linux command that allows you to look at a file. | ||
*<code>-S</code> option prevents line wrap. | *<code>-S</code> option prevents line wrap. | ||
| Line 95: | Line 95: | ||
** 1000G_omni2.5.b37.sites.PASS.chr7.vcf.gz.tbi | ** 1000G_omni2.5.b37.sites.PASS.chr7.vcf.gz.tbi | ||
** Used for variant filtering | ** Used for variant filtering | ||
| + | |||
| + | * INDEL sites | ||
| + | ** 1kg.pilot_release.merged.indels.sites.hg19.chr7.vcf | ||
| + | ** Used for variant calling | ||
==== GotCloud FASTQ Index File ==== | ==== GotCloud FASTQ Index File ==== | ||
| Line 164: | Line 168: | ||
Sample2 fastq/S2/F2.fastq.gz . RGID2 SampleID2 Lib2 UM ILLUMINA | Sample2 fastq/S2/F2.fastq.gz . RGID2 SampleID2 Lib2 UM ILLUMINA | ||
| − | The <code>--fastq</code> | + | The command-line <code>--fastq</code> option or the configuration file <code>FASTQ_PREFIX</code> setting can be used to specify a prefix to the FASTQ1/FASTQ2 file paths. |
| − | + | This file is specified either via the command-line <code>--index_file</code> parameter or via the configuration file <code>INDEX_FILE</code> setting. | |
| − | This file is specified either via the command line | ||
The command-line setting takes precedence over the configuration file setting. | The command-line setting takes precedence over the configuration file setting. | ||
| Line 173: | Line 176: | ||
==== GotCloud Configuration File ==== | ==== GotCloud Configuration File ==== | ||
This file is created by you to configure GotCloud for your data. | This file is created by you to configure GotCloud for your data. | ||
| + | |||
| + | * Default values are provided in ${GC}/gotcloud/bin/gotcloudDefaults.conf | ||
| + | ** Most values should be left as the defaults | ||
| + | * Specify values in your configuration file as: | ||
| + | KEY = value | ||
| + | * Keys to override: | ||
| + | {|class="wikitable" cellpadding=5 | ||
| + | |- | ||
| + | ! Key Name !! Description | ||
| + | |- | ||
| + | | colspan=2 style="text-align:center"| Index File Settings - pointing GotCloud to your data | ||
| + | |- | ||
| + | | INDEX_FILE || Path to the FASTQ index file that you created | ||
| + | * Alternatively, this can be specified on the command-line as <code>--index_file</code> | ||
| + | |- | ||
| + | | FASTQ_PREFIX || Prefix to be added to the FASTQ files in INDEX_FILE | ||
| + | * Not required | ||
| + | |- | ||
| + | | BAM_INDEX || Path to the BAM index file | ||
| + | * to be created by alignment | ||
| + | * to be used for snp calling | ||
| + | |- | ||
| + | | colspan=2 style="text-align:center"| Reference File Settings - telling GotCloud where to find your reference files | ||
| + | |- | ||
| + | | REF_DIR || Path to your reference files | ||
| + | * You don't have to use this, you can specify the full path for each file | ||
| + | |- | ||
| + | | REF || Path/filename of the FASTA reference file | ||
| + | * If different than default: $(REF_DIR)/human.g1k.v37.fa | ||
| + | |- | ||
| + | | DBSNP_VCF || Path/filename of the DBSNP file | ||
| + | * If different than default: $(REF_DIR)/dbsnp_135.b37.vcf.gz | ||
| + | |- | ||
| + | | HM3_VCF || Path/filename of the HapMap3 file | ||
| + | * If different than default: $(REF_DIR)/hapmap_3.3.b37.sites.vcf.gz | ||
| + | |- | ||
| + | | OMNI_VCF || Path/filename of the OMNI file | ||
| + | * If different than default: $(REF_DIR)/1000G_omni2.5.b37.sites.PASS.vcf.gz | ||
| + | |- | ||
| + | | INDEL_PREFIX || Path/filename base of the indels file | ||
| + | * If different than default: $(REF_DIR)/1kg.pilot_release.merged.indels.sites.hg19 | ||
| + | |} | ||
Revision as of 11:59, 11 June 2014
Step 0: Login to the machine & setup environment
- Login to the windows machine
- The username/password for the Windows machine should be written on it
- Open putty
- Start->.....
- In putty, login to seqshop-server.sph.umich.edu
- Server name: seqshop-server.sph.umich.edu
- Enter your provided username & password
- To simplify commands/typing, we will setup an environment variable to point to the GotCloud directory.
export GC=/home/mktrost/seqshop/
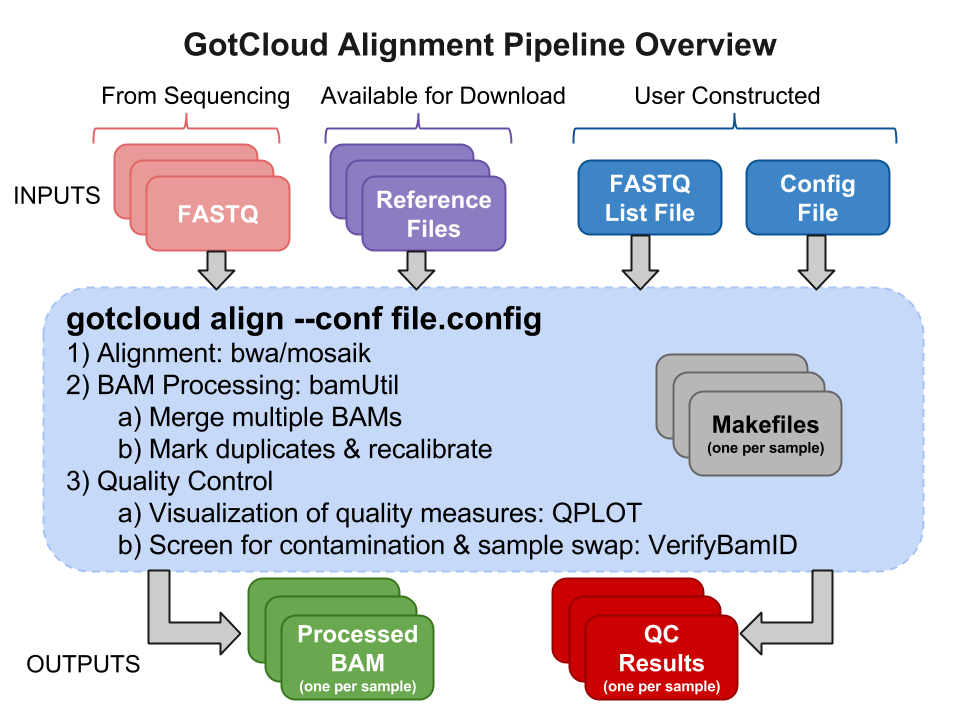
GotCloud Alignment Pipeline
Input Files
Sequence Data Files : FASTQs
The FASTQ files are provided to you by those who did the sequencing.
For this tutorial, we will use FASTQs for 6 1000Genome samples
ls ${GC}/inputs/fastq/
There are 51 fastq files: combination of single-end & paired-end.
- Single-end: HG00641.chr7.CFTR.SRR069531.fastq
- Paired-end: HG00641.chr7.CFTR.SRR069531_1.fastq & HG00641.chr7.CFTR.SRR069531_2.fastq
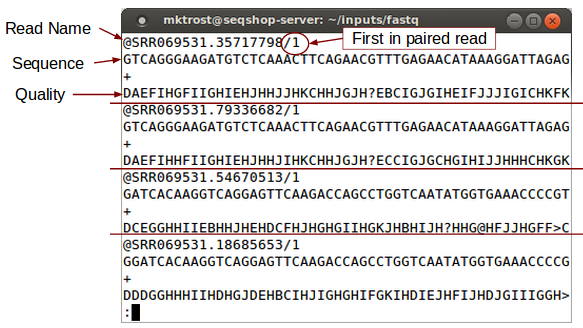
Look at FASTQ:
less -S ${GC}/inputs/fastq/HG00641.chr7.CFTR.SRR069531_1.fastq
less is a Linux command that allows you to look at a file.
-Soption prevents line wrap.- Use the arrow (up/down/left/right) keys to scroll through the file.
- Use
zlessif the file is compressed.
Reference Files
Reference files can be downloaded with GotCloud or from other sources.
ls ${GC}/reference/chr7
Reference FASTA File (All reference bases for chromosome)
- human.g1k.v37.chr7.fa
- human.g1k.v37.chr7.fa.fai - index for .fa file
Let's look at the reference file:
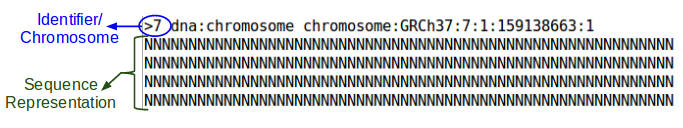
head -n 5 ${GC}/reference/chr7/human.g1k.v37.chr7.fa
All N's, so let's look at a later section:
tail -n+2000 ${GC}/reference/chr7/human.g1k.v37.chr7.fa |head -n 5
Our binary representation of the reference file:
- human.g1k.v37.chr7-bs.umfa
- Automatically generated by our tools if it doesn't exist.
BWA Aligner Specific Reference Files
- human.g1k.v37.chr7.fa.bwt
- human.g1k.v37.chr7.fa.pac
- human.g1k.v37.chr7.fa.ann
- human.g1k.v37.chr7.fa.amb
- human.g1k.v37.chr7.fa.sa
Mosaik Aligner Specific Reference Files
- pe.100.01.ann
- se.100.005.ann
- human.g1k.v37.chr7.dat
- human.g1k.v37.chr7_15_meta.jmp
- human.g1k.v37.chr7_15_keys.jmp
- human.g1k.v37.chr7_15_positions.jmp
QPLOT Reference File
- human.g1k.v37.chr7.winsize100.gc
- QPLOT can create if needed
Variant Files
- Known variants in DBSNP:
- dbsnp_135.b37.chr7.vcf.gz
- dbsnp_135.b37.chr7.vcf.gz.tbi
- Used to skip known variant sites for recalibration
- Used for variant filtering
- List of hapmap sites
- hapmap_3.3.b37.sites.chr7.vcf.gz
- hapmap_3.3.b37.sites.chr7.vcf.gz.tbi
- Used for contamination/sample swap validation
- used for variant filtering
- OMNI sites.
- 1000G_omni2.5.b37.sites.PASS.chr7.vcf.gz
- 1000G_omni2.5.b37.sites.PASS.chr7.vcf.gz.tbi
- Used for variant filtering
- INDEL sites
- 1kg.pilot_release.merged.indels.sites.hg19.chr7.vcf
- Used for variant calling
GotCloud FASTQ Index File
This file is created by you and directs GotCloud to your FASTQ files, providing additional information for them.
- tab delimited
- columns may be in any order
- starts with a header line
- one line per single-end read
- one line per paired-end read (only 1 line per pair).
Required Columns
| Column Name | Description | Recommended Value |
|---|---|---|
| MERGE_NAME |
|
Sample Name |
| FASTQ1 |
|
path/fastq1 |
| FASTQ2 |
|
path/fastq2 |
The following columns are optional and used to populate the Read Group Information in the BAM file.
- RGID field is required if using any of these fields, the others are optional.
What is a Read Group?
- Groups reads together
- Used for recalibration
- Each sequencing run should get a different ReadGroup
If you do not want the field for:
- any fastq, leave the column out of the header line
- a single line, use a '.'
| Column Name | Description | Recommended Value |
|---|---|---|
| RGID | Read Group ID | Run ID |
| SAMPLE | Sample Name | Sample Name |
| LIBRARY | Library
|
if you don't know or it is all the same, use Sample Name |
| CENTER | Center Name | Name of the sequencing center producing the FASTQ |
| PLATFORM | Platform | CAPILLARY, LS454, ILLUMINA,
SOLID, HELICOS, IONTORRENT, or PACBIO |
MERGE_NAME FASTQ1 FASTQ2 RGID SAMPLE LIBRARY CENTER PLATFORM Sample1 fastq/S1/F1_R1.fastq.gz fastq/S1/F1_R2.fastq.gz RGID1 SampleID1 Lib1 UM ILLUMINA Sample1 fastq/S1/F2_R1.fastq.gz fastq/S1/F2_R2.fastq.gz RGID1a SampleID1 Lib1 UM ILLUMINA Sample2 fastq/S2/F1_R1.fastq.gz fastq/S2/F1_R2.fastq.gz RGID2 SampleID2 Lib2 UM ILLUMINA Sample2 fastq/S2/F2.fastq.gz . RGID2 SampleID2 Lib2 UM ILLUMINA
The command-line --fastq option or the configuration file FASTQ_PREFIX setting can be used to specify a prefix to the FASTQ1/FASTQ2 file paths.
This file is specified either via the command-line --index_file parameter or via the configuration file INDEX_FILE setting.
The command-line setting takes precedence over the configuration file setting.
GotCloud Configuration File
This file is created by you to configure GotCloud for your data.
- Default values are provided in ${GC}/gotcloud/bin/gotcloudDefaults.conf
- Most values should be left as the defaults
- Specify values in your configuration file as:
KEY = value
- Keys to override:
| Key Name | Description |
|---|---|
| Index File Settings - pointing GotCloud to your data | |
| INDEX_FILE | Path to the FASTQ index file that you created
|
| FASTQ_PREFIX | Prefix to be added to the FASTQ files in INDEX_FILE
|
| BAM_INDEX | Path to the BAM index file
|
| Reference File Settings - telling GotCloud where to find your reference files | |
| REF_DIR | Path to your reference files
|
| REF | Path/filename of the FASTA reference file
|
| DBSNP_VCF | Path/filename of the DBSNP file
|
| HM3_VCF | Path/filename of the HapMap3 file
|
| OMNI_VCF | Path/filename of the OMNI file
|
| INDEL_PREFIX | Path/filename base of the indels file
|