Difference between revisions of "Test EPACTS for DIAGRAM"
Clement Ma (talk | contribs) |
|||
| (14 intermediate revisions by one other user not shown) | |||
| Line 1: | Line 1: | ||
= Download EPACTS = | = Download EPACTS = | ||
| − | EPACTS is available for download [http:// | + | EPACTS is available for download [http://csg.sph.umich.edu//kang/epacts/download/epacts_v2.12.noref_binary.2012_10_01.tar.gz here (100Mb) ]. |
Requirements | Requirements | ||
| Line 11: | Line 11: | ||
Uncompress EPACTS package to the directory you would like to install | Uncompress EPACTS package to the directory you would like to install | ||
| − | + | <pre> tar xzvf epacts_v2_12.noref_binary.2012_10_01.tar.gz</pre> | |
| − | + | Download the reference FASTA files from 1000 Genomes FTP automatically by running the following commands<br> | |
| + | <pre>cd epacts2.1/ | ||
| + | ./ref_download.sh | ||
| + | (For advanced users, to save time for downloading the FASTA files (~900MB), you may copy a local copy of GRCh37 FASTA file and the index file to ${EPACTS_DIR}/ext/ref/) | ||
| + | </pre> | ||
= Accessing help = | = Accessing help = | ||
For a list of commands available in EPACTS, type in the following commands: | For a list of commands available in EPACTS, type in the following commands: | ||
| − | <pre>$ epacts2/epacts help | + | <pre>$ epacts2.1/epacts help |
| + | |||
Usage: | Usage: | ||
epacts [command] [options] | epacts [command] [options] | ||
| Line 29: | Line 34: | ||
zoom Create a locus zoom plot from epacts results | zoom Create a locus zoom plot from epacts results | ||
meta Perform meta-analysis across multiple epacts results | meta Perform meta-analysis across multiple epacts results | ||
| + | make-group Create the group information for gene-based testing | ||
| + | make-kin Create a kinship matrix | ||
Visit http://genome.sph.umich.edu/wiki/EPACTS for more detailed documentation | Visit http://genome.sph.umich.edu/wiki/EPACTS for more detailed documentation | ||
| − | </pre> | + | |
| − | + | </pre> | |
| − | <pre>$ epacts2/epacts single -help | + | |
| + | |||
| + | To view options for single variant testing only type in: | ||
| + | <pre>$ epacts2.1/epacts single -help | ||
Usage: | Usage: | ||
epacts single [options] | epacts single [options] | ||
| Line 45: | Line 55: | ||
</pre> | </pre> | ||
| − | = Getting started in EPACTS with an example = | + | = Getting started in EPACTS with an example = |
Once installed, test out the software by running a quick example using the test data provided in the "example" directory. The example VCF and PED files are: | Once installed, test out the software by running a quick example using the test data provided in the "example" directory. The example VCF and PED files are: | ||
| − | <pre>$ epacts2/example/1000G_exome_chr20_example_softFiltered.calls.vcf.gz | + | <pre>$ epacts2.1/example/1000G_exome_chr20_example_softFiltered.calls.vcf.gz |
| − | $ epacts2/example/1000G_dummy_pheno.ped | + | $ epacts2.1/example/1000G_dummy_pheno.ped |
</pre> | </pre> | ||
<br> Run the single variant score test on the example data using this command: | <br> Run the single variant score test on the example data using this command: | ||
| − | <pre>$ epacts2/epacts single \ | + | <pre>$ epacts2.1/epacts single \ |
| − | --vcf epacts2/example/1000G_exome_chr20_example_softFiltered.calls.vcf.gz \ | + | --vcf epacts2.1/example/1000G_exome_chr20_example_softFiltered.calls.vcf.gz \ |
| − | --ped epacts2/example/1000G_dummy_pheno.ped \ | + | --ped epacts2.1/example/1000G_dummy_pheno.ped \ |
--min-maf 0.001 --chr 20 --pheno DISEASE --cov AGE --cov SEX --test b.score --anno \ | --min-maf 0.001 --chr 20 --pheno DISEASE --cov AGE --cov SEX --test b.score --anno \ | ||
--out {OUTPUT_DIR}/test --run 2 & | --out {OUTPUT_DIR}/test --run 2 & | ||
| Line 66: | Line 76: | ||
1. '''test.epacts.gz''' contains all the association results. | 1. '''test.epacts.gz''' contains all the association results. | ||
| − | <pre> | + | <pre>$ head {OUTPUT_DIR}/test.epacts |
| − | #CHROM BEGIN END MARKER_ID NS AC CALLRATE MAF PVALUE SCORE | + | #CHROM BEGIN END MARKER_ID NS AC CALLRATE MAF PVALUE SCORE N.CASE N.CTRL AF.CASE AF.CTRL |
| − | 20 68303 68303 20:68303_A/G_Upstream:DEFB125 266 1 1 0.0018797 NA NA | + | 20 68303 68303 20:68303_A/G_Upstream:DEFB125 266 1 1 0.0018797 NA NA NA NA NA NA |
| − | 20 68319 68319 20:68319_C/A_Upstream:DEFB125 266 0 1 0 NA NA | + | 20 68319 68319 20:68319_C/A_Upstream:DEFB125 266 0 1 0 NA NA NA NA NA NA |
| − | 20 68396 68396 20:68396_C/T_Nonsynonymous:DEFB125 266 1 1 0.0018797 NA NA | + | 20 68396 68396 20:68396_C/T_Nonsynonymous:DEFB125 266 1 1 0.0018797 NA NA NA NA NA NA |
| − | 20 76635 76635 20:76635_A/T_Intron:DEFB125 266 0 1 0 NA NA | + | 20 76635 76635 20:76635_A/T_Intron:DEFB125 266 0 1 0 NA NA NA NA NA NA |
| − | 20 76689 76689 20:76689_T/C_Synonymous:DEFB125 266 0 1 0 NA NA | + | 20 76689 76689 20:76689_T/C_Synonymous:DEFB125 266 0 1 0 NA NA NA NA NA NA |
| − | 20 76690 76690 20:76690_T/C_Nonsynonymous:DEFB125 266 1 1 0.0018797 NA NA | + | 20 76690 76690 20:76690_T/C_Nonsynonymous:DEFB125 266 1 1 0.0018797 NA NA NA NA NA NA |
| − | 20 76700 76700 20:76700_G/A_Nonsynonymous:DEFB125 266 0 1 0 NA NA | + | 20 76700 76700 20:76700_G/A_Nonsynonymous:DEFB125 266 0 1 0 NA NA NA NA NA NA |
| − | 20 76726 76726 20:76726_C/G_Nonsynonymous:DEFB125 266 0 1 0 NA NA | + | 20 76726 76726 20:76726_C/G_Nonsynonymous:DEFB125 266 0 1 0 NA NA NA NA NA NA |
| − | 20 76771 76771 20:76771_C/T_Nonsynonymous:DEFB125 266 3 1 0.0056391 0.68484 0.40587 | + | 20 76771 76771 20:76771_C/T_Nonsynonymous:DEFB125 266 3 1 0.0056391 0.68484 0.40587 145 121 0.013793 0.0082645 |
</pre> | </pre> | ||
| + | The columns in the results file are: | ||
| + | |||
| + | #CHROM: chromosome | ||
| + | #BEGIN: starting position | ||
| + | #END: ending position (same as BEGIN if a SNP) | ||
| + | #MARKER_ID: name of varian | ||
| + | #NS: Number of samples (cases + controls) | ||
| + | #AC: Total allele count in sample | ||
| + | #CALLRATE: call rate | ||
| + | #MAF: minor allele frequency in full sample | ||
| + | #PVALUE: score test association p-value | ||
| + | #SCORE: test statistic for score test | ||
| + | #N.CASE: number of cases | ||
| + | #N.CTRL: number of controls | ||
| + | #AF.CASE: allele frequency in cases only | ||
| + | #AF.CTRL: allele frequency in controls only | ||
| + | |||
| + | Note: For variants below the minimum MAF threshold (min-maf = 0.001), the number of cases and controls (N.CASE, N.CTRL) are not outputted (listed as "NA"). | ||
| + | |||
| + | <br> | ||
| + | |||
2. '''test.epacts.top5000''' contains the top 5000 associated variants ordered by p-value. | 2. '''test.epacts.top5000''' contains the top 5000 associated variants ordered by p-value. | ||
| − | <pre>$ head | + | <pre>$ head {OUTPUT_DIR}/test.epacts.top5000 |
| − | #CHROM BEGIN END MARKER_ID NS AC CALLRATE MAF PVALUE SCORE | + | #CHROM BEGIN END MARKER_ID NS AC CALLRATE MAF PVALUE SCORE N.CASE N.CTRL AF.CASE AF.CTRL |
| − | 20 1610894 1610894 20:1610894_G/A_Synonymous:SIRPG 266 136 1 0.25564 0.0001097 3.8681 | + | 20 1610894 1610894 20:1610894_G/A_Synonymous:SIRPG 266 136 1 0.25564 0.0001097 3.8681 145 121 0.64138 0.35537 |
| − | 20 4162411 4162411 20:4162411_T/C_Intron:SMOX 266 204 1 0.38346 0.00055585 -3.4523 | + | 20 4162411 4162411 20:4162411_T/C_Intron:SMOX 266 204 1 0.38346 0.00055585 -3.4523 145 121 0.62759 0.93388 |
| − | 20 34061918 34061918 20:34061918_T/C_Intron:CEP250 266 39 1 0.073308 0.0011231 3.2577 | + | 20 34061918 34061918 20:34061918_T/C_Intron:CEP250 266 39 1 0.073308 0.0011231 3.2577 145 121 0.21379 0.066116 |
| − | 20 4155948 4155948 20:4155948_G/A_Intron:SMOX 266 215 1 0.40414 0.0020791 -3.0787 | + | 20 4155948 4155948 20:4155948_G/A_Intron:SMOX 266 215 1 0.40414 0.0020791 -3.0787 145 121 0.68276 0.95868 |
| − | 20 4680251 4680251 20:4680251_A/G_Nonsynonymous:PRNP 266 186 1 0.34962 0.0025962 3.0119 | + | 20 4680251 4680251 20:4680251_A/G_Nonsynonymous:PRNP 266 186 1 0.34962 0.0025962 3.0119 145 121 0.8069 0.57025 |
| − | 20 36668874 36668874 20:36668874_G/A_Synonymous:RPRD1B 266 96 1 0.18045 0.003031 2.9646 | + | 20 36668874 36668874 20:36668874_G/A_Synonymous:RPRD1B 266 96 1 0.18045 0.003031 2.9646 145 121 0.44828 0.2562 |
| − | 20 36641871 36641871 20:36641871_G/A_Synonymous:TTI1 266 10 1 0.018797 0.004308 -2.8547 | + | 20 36641871 36641871 20:36641871_G/A_Synonymous:TTI1 266 10 1 0.018797 0.004308 -2.8547 145 121 0.0068966 0.07438 |
| − | 20 32664926 32664926 20:32664926_G/A_Nonsynonymous:RALY 266 20 1 0.037594 0.0046365 2.8313 | + | 20 32664926 32664926 20:32664926_G/A_Nonsynonymous:RALY 266 20 1 0.037594 0.0046365 2.8313 145 121 0.11724 0.024793 |
| − | 20 34288854 34288854 20:34288854_C/T_Utr3:ROMO1 266 28 1 0.052632 0.0047722 2.822 | + | 20 34288854 34288854 20:34288854_C/T_Utr3:ROMO1 266 28 1 0.052632 0.0047722 2.822 145 121 0.15862 0.041322 |
| + | |||
</pre> | </pre> | ||
3. '''test.epacts.qq.pdf''' contains the Q-Q plot of test p-values (stratified by MAF)<br>[[Image:Test b score epacts qq.png]] | 3. '''test.epacts.qq.pdf''' contains the Q-Q plot of test p-values (stratified by MAF)<br>[[Image:Test b score epacts qq.png]] | ||
Latest revision as of 10:28, 2 February 2017
Download EPACTS
EPACTS is available for download here (100Mb) .
Requirements
- Linux 64bit
- Perl 5
Install EPACTS
Uncompress EPACTS package to the directory you would like to install
tar xzvf epacts_v2_12.noref_binary.2012_10_01.tar.gz
Download the reference FASTA files from 1000 Genomes FTP automatically by running the following commands
cd epacts2.1/
./ref_download.sh
(For advanced users, to save time for downloading the FASTA files (~900MB), you may copy a local copy of GRCh37 FASTA file and the index file to ${EPACTS_DIR}/ext/ref/)
Accessing help
For a list of commands available in EPACTS, type in the following commands:
$ epacts2.1/epacts help Usage: epacts [command] [options] Command: help Print out brief help message man Print the full documentation in man page style single Perform single variant association group Perform groupwise (burden-style) association test anno Annotate a VCF file zoom Create a locus zoom plot from epacts results meta Perform meta-analysis across multiple epacts results make-group Create the group information for gene-based testing make-kin Create a kinship matrix Visit http://genome.sph.umich.edu/wiki/EPACTS for more detailed documentation
To view options for single variant testing only type in:
$ epacts2.1/epacts single -help Usage: epacts single [options] Required Options (Run epacts single -man or see wiki for more info): -vcf STR Input VCF file (tabixed and bgzipped) -ped STR Input PED file for phenotypes and covariates -out STR Prefix of output files -test STR Statistical test to use ...
Getting started in EPACTS with an example
Once installed, test out the software by running a quick example using the test data provided in the "example" directory. The example VCF and PED files are:
$ epacts2.1/example/1000G_exome_chr20_example_softFiltered.calls.vcf.gz $ epacts2.1/example/1000G_dummy_pheno.ped
Run the single variant score test on the example data using this command:
$ epacts2.1/epacts single \
--vcf epacts2.1/example/1000G_exome_chr20_example_softFiltered.calls.vcf.gz \
--ped epacts2.1/example/1000G_dummy_pheno.ped \
--min-maf 0.001 --chr 20 --pheno DISEASE --cov AGE --cov SEX --test b.score --anno \
--out {OUTPUT_DIR}/test --run 2 &
This command will run the single variant test on the input VCF and PED files, with a minimum MAF threshold of 0.001. The phenotype is "DISEASE" and we are adjusting the analysis with covariates AGE and SEX. The output file directory prefix is {OUTPUT_DIR}/test. Finally, EPACTS will run the analysis in parallel on 2 CPUs.
Expected output
EPACTS produces a number of files and plots.
1. test.epacts.gz contains all the association results.
$ head {OUTPUT_DIR}/test.epacts
#CHROM BEGIN END MARKER_ID NS AC CALLRATE MAF PVALUE SCORE N.CASE N.CTRL AF.CASE AF.CTRL
20 68303 68303 20:68303_A/G_Upstream:DEFB125 266 1 1 0.0018797 NA NA NA NA NA NA
20 68319 68319 20:68319_C/A_Upstream:DEFB125 266 0 1 0 NA NA NA NA NA NA
20 68396 68396 20:68396_C/T_Nonsynonymous:DEFB125 266 1 1 0.0018797 NA NA NA NA NA NA
20 76635 76635 20:76635_A/T_Intron:DEFB125 266 0 1 0 NA NA NA NA NA NA
20 76689 76689 20:76689_T/C_Synonymous:DEFB125 266 0 1 0 NA NA NA NA NA NA
20 76690 76690 20:76690_T/C_Nonsynonymous:DEFB125 266 1 1 0.0018797 NA NA NA NA NA NA
20 76700 76700 20:76700_G/A_Nonsynonymous:DEFB125 266 0 1 0 NA NA NA NA NA NA
20 76726 76726 20:76726_C/G_Nonsynonymous:DEFB125 266 0 1 0 NA NA NA NA NA NA
20 76771 76771 20:76771_C/T_Nonsynonymous:DEFB125 266 3 1 0.0056391 0.68484 0.40587 145 121 0.013793 0.0082645
The columns in the results file are:
- CHROM: chromosome
- BEGIN: starting position
- END: ending position (same as BEGIN if a SNP)
- MARKER_ID: name of varian
- NS: Number of samples (cases + controls)
- AC: Total allele count in sample
- CALLRATE: call rate
- MAF: minor allele frequency in full sample
- PVALUE: score test association p-value
- SCORE: test statistic for score test
- N.CASE: number of cases
- N.CTRL: number of controls
- AF.CASE: allele frequency in cases only
- AF.CTRL: allele frequency in controls only
Note: For variants below the minimum MAF threshold (min-maf = 0.001), the number of cases and controls (N.CASE, N.CTRL) are not outputted (listed as "NA").
2. test.epacts.top5000 contains the top 5000 associated variants ordered by p-value.
$ head {OUTPUT_DIR}/test.epacts.top5000
#CHROM BEGIN END MARKER_ID NS AC CALLRATE MAF PVALUE SCORE N.CASE N.CTRL AF.CASE AF.CTRL
20 1610894 1610894 20:1610894_G/A_Synonymous:SIRPG 266 136 1 0.25564 0.0001097 3.8681 145 121 0.64138 0.35537
20 4162411 4162411 20:4162411_T/C_Intron:SMOX 266 204 1 0.38346 0.00055585 -3.4523 145 121 0.62759 0.93388
20 34061918 34061918 20:34061918_T/C_Intron:CEP250 266 39 1 0.073308 0.0011231 3.2577 145 121 0.21379 0.066116
20 4155948 4155948 20:4155948_G/A_Intron:SMOX 266 215 1 0.40414 0.0020791 -3.0787 145 121 0.68276 0.95868
20 4680251 4680251 20:4680251_A/G_Nonsynonymous:PRNP 266 186 1 0.34962 0.0025962 3.0119 145 121 0.8069 0.57025
20 36668874 36668874 20:36668874_G/A_Synonymous:RPRD1B 266 96 1 0.18045 0.003031 2.9646 145 121 0.44828 0.2562
20 36641871 36641871 20:36641871_G/A_Synonymous:TTI1 266 10 1 0.018797 0.004308 -2.8547 145 121 0.0068966 0.07438
20 32664926 32664926 20:32664926_G/A_Nonsynonymous:RALY 266 20 1 0.037594 0.0046365 2.8313 145 121 0.11724 0.024793
20 34288854 34288854 20:34288854_C/T_Utr3:ROMO1 266 28 1 0.052632 0.0047722 2.822 145 121 0.15862 0.041322
3. test.epacts.qq.pdf contains the Q-Q plot of test p-values (stratified by MAF)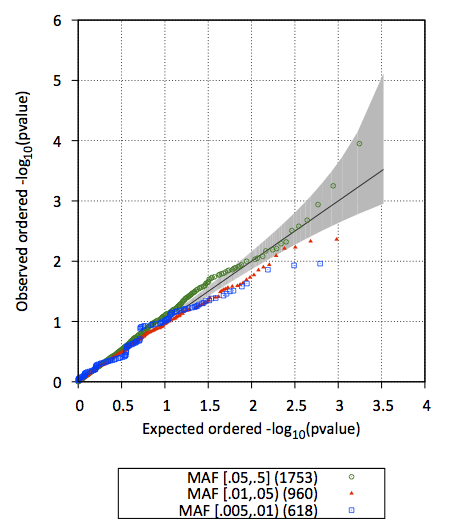
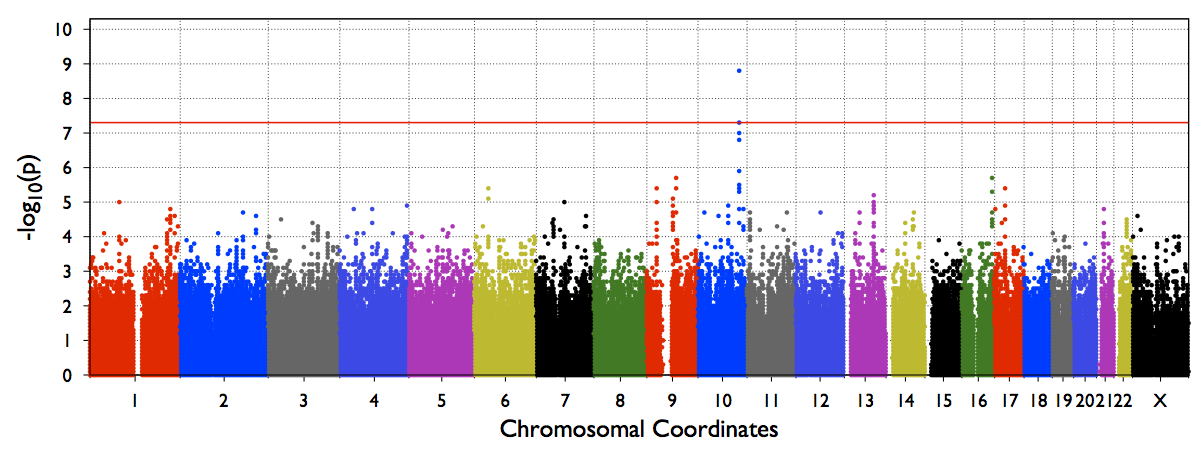
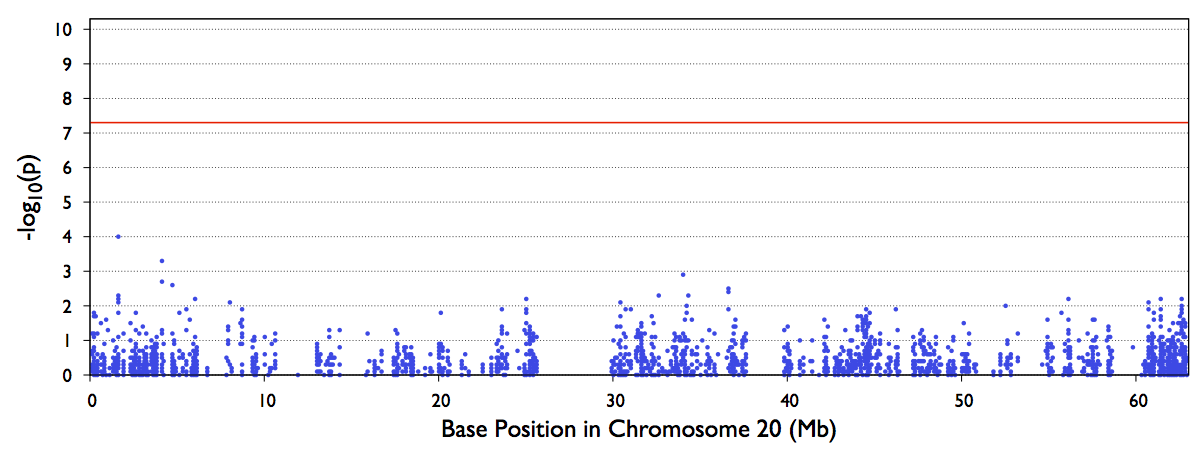
4. test.epacts.mh.pdf contains the Manhattan Plot of test p-values
The file out/test.b.score.epacts.mh.pdf will be generated for chr20 only.
An example Genome-wide manhattan plot (from a genome-wide run) will look like below