EPACTS
EPACTS (Efficient and Parallelizable Association Container Toolbox) is a versatile software pipeline to perform various statistical tests for identifying genome-wide association from sequence data through a user-friendly interface, both to scientific analysts and to method developers.
Join in EPACTS mailing list
Please join in the EPACTS Google Group to ask / discuss / comment about EPACTS.
Lastest ChangeLog
- Dec 15th, 2016 : EPACTS v3.3.0 release (github)
- Moved the repository into github
- Some major fixes in handling large sample size (>18,000)
- Other minor bug fixes
- July 10th, 2014 : EPACTS v3.2.6 release
- Minor bug fix in epacts-make-kin
- March 11th, 2014 : EPACTS v3.2.5 release
- EMMAX-SKAT is implemented with major bug fix
- November 21th, 2013 : EPACTS v3.2.4 release
- Fixed a number of minor bugs (more comprehensive fix is still pending)
- March 25th, 2013 : EPACTS v3.2.3 release
- Relaxed the checking of low-rank matrix in SKAT tests (to avoid unncessary skipping of genes)
- March 13th, 2013 : EPACTS v3.2.2 release
- Fixed an error which occasionally report mismatches in the number of samples
- March 9th, 2013 : EPACTS v3.2.1 release
- Fixed errors in loading the dynamic library
- Fixed errors in SKAT-O (thanks to Anubha Mahajan and Jason Flannick)
- Fixed bugs in emmax-CMC
- Added emmax-SKAT (contributed by Seunngeun Lee)
- And additional minor bug fixes
See #Full ChangeLog for full details
Key Features
EPACTS currently provides the following set of key features
- Robust support for widely used format of sequence-based genotypes (VCF) and phenotypes with pedigree (PED)
- Efficient library for accessing VCF file to reduce computational burden to analyze large-scale sequencing data
- Support selecting markers by arbitrary combination of substring matching.
- Support for using genotype dosages instead of hard genotype calls
- Utilize PED format to perform test across multiple traits.
- Supports a large number of widely used statistical tests for single variant association and burden tests.
- See the "Currently Supported Statistical Tests" section below for more information
- Easy to Highly Parallelize Jobs
- Makefile-based partition into and ligation of multiple subtasks
- Parallel run of job is simply adding one parameter when running EPACTS
- Integrative and versatile framework that allows easy addition of additional statistical test
- Core input/output routines are implemented in C++
- Most statistical tests (except for EMMAX) are implemented in R
- Adding a simple R function to implement additional statistical test (See #Implementing Additional Statistical Tests for details)
- Useful utilities for post-association-analysis tasks
- Automatic functional annotation of associated variants
- Automatic generation of QQ and Manhattan Plot
- (TBA) Zoom plot for the significant associations
Obtaining EPACTS
- The official release of EPACTS software is available at https://github.com/statgen/EPACTS
- From the CSG cluster, it is available at /net/fantasia/home/bin/epacts/
- Note that R (version 2.10 or higher) and gnuplot (version 4.2 or higher) must be installed in order to run EPACTS correctly.
Currently Supported Statistical Tests
EPACTS supports the following sets of widely used statistical tests for single variant tests and burden tests
Single Variant Tests
| Test Name | Phenotypes | Covariates | Computational Time | Description | Implemented by |
|---|---|---|---|---|---|
| b.wald | Binary | YES (Joint) |
Slow | Logisitic Wald Test | Hyun Min Kang (simply used glm in R) |
| b.score | Binary | YES (Regressed Out) |
Fast | Logistic Score Test (from Lin DY and Tang ZZ, AJHG 2011 89:354-67) |
Clement Ma & Hyun Min Kang |
| b.firth | Binary | YES (Joint) |
Slow | Firth Bias-Corrected Logistic Likelihood Ratio Test | Clement Ma |
| b.spa2 | Binary | YES |
Moderate | Saddlepoint Approximation Method | Shawn Lee & Rounak Dey |
| b.lrt | Binary | YES (Joint) |
Slow | Likelihood Ratio Test | Clement Ma |
| b.glrt | Binary | NO | Fast | Genotype Likelihood Ratio Test (use GL or PL field in VCF to perform case-control test) |
Hyun Min Kang |
| q.lm | Quantitative | YES (Joint) |
Slow | Linear Wald Test | Hyun Min Kang (as implemented in lm in R) |
| q.linear | Quantitative | YES (Regressed Out) |
Fast | Linear Wald Test | Hyun Min Kang |
| q.reverse | Quantitative | YES (Joint) |
Slow | Reverse regression of phenotypes on binary genotypes (dominant model) |
Hyun Min Kang |
| q.wilcox | Quantitative | YES (Regressed Out First) |
Slow | Nonparametric Reverse regression of phenotypes on binary genotypes (dominant model) |
Hyun Min Kang |
| q.emmax | Quantitative | YES (Regressed Out First) |
Slow | EMMAX ( Kang et al (2010) Nat Genet 42:348-54 ) |
Hyun Min Kang |
Gene-wise or group-wise tests
| Test Name | Phenotypes | Covariates | Computational Time | Description | Implemented by |
|---|---|---|---|---|---|
| b.collapse | Binary | YES (Joint Estimation) |
Slow | Logistic Wald Test between binary phenotypes and 0/1 collapsed variables | Hyun Min Kang |
| b.madsen | Binary | NO | Slow | Wilcoxon Rank Sum Test between binary phenotypes and weighted rare variant scores (slightly different version from the published method - it uses pooled allele frequency across cases and controls for weighting each variant) | Hyun Min Kang |
| b.wcnt | Binary | YES (Joint Estimation) |
Slow | Logistic Wald Test between binary phenotypes and weighted rare variant scores | Hyun Min Kang |
| q.reverse | Quantitative | YES (Joint Estimation) |
Slow | Reverse regression of phenotypes on binary collapsed variables | Hyun Min Kang |
| q.wilcox | Quantitative | YES (Regressed Out First) |
Slow | Nonparametric Reverse regression of phenotypes on collapsed variables | Hyun Min Kang |
| skat | Binary/Quantitative | YES (Joint Estimation) |
Slow | SKAT-O Test by Lee et al, Biostatistics (2012) | Seunggeun Lee (adaptive by Xueling Sim and Hyun Min Kang) |
| VT | Binary/Quantitative | YES (Regressed out first) |
Slow | Variable Threshold Test with adaptive permutation Price et al, AJHG (2010) 86:832-8 |
Hyun Min Kang |
| emmaxCMC | Binary/Quantitative | YES (Regressed Out First) |
Slow | Collapsing burden test using EMMAX | Hyun Min Kang |
| emmaxVT | Binary/Quantitative | YES (Regressed Out First) |
Slow | Variable-threshold burden test using EMMAX | Hyun Min Kang |
| mmskat | Quantitative | YES (Regressed Out First) |
Slow | SKAT test using EMMAX | Seunggeun Lee & Hyun Min Kang |
Installation Details
If you want to use EPACTS in an Ubuntu platform, following the step below
$ git clone https://github.com/statgen/EPACTS.git $ cd EPACTS $ ./configure --prefix [/path/to/install] $ make $ make install
(Important Note: make sure to specify --prefix=/path/to/install to avoid installing to the default path /usr/local/, which you may not have the permission. /home/your_userid/epacts might be a good one, if you are not sure where to install)
- Now ${EPACTS_DIR} represents the '/path/to/install' directory
- Download the reference FASTA files from 1000 Genomes FTP automatically by running the following commands
${EPACTS_DIR}/bin/epacts download
(For advanced users, to save time for downloading the FASTA files (~900MB), you may copy a local copy of GRCh37 FASTA file and the index file to ${EPACTS_DIR}/share/EPACTS/)
- Perform a test run by running the following command
${EPACTS_DIR}/bin/test_run_epacts.sh
In order to use EPACTS in the CSG cluster, you do not need to install them. You can directly use or make a copy of the in-house release version at
/net/fantasia/home/hmkang/tools/epacts-3.3.0/bin/epacts/
- If you want to access previous versions, visit http://csg-old.sph.umich.edu/kang/epacts/download
Getting Started With Examples
If you are using EPACTS from the CSG cluster, please set the following environment variable
EPACTS_DIR=/net/fantasia/home/hmkang/tools/epacts-3.3.0/bin/epacts (in bash) setenv EPACTS_DIR /net/fantasia/home/hmkang/tools/epacts-3.3.0/bin/epacts (in csh)
If you downloaded EPACTS binary and please set EPACTS_DIR to the full path of the downloaded and uncompressed directory.
All-in-one example
To get started with EPACTS, run the following command will perform an example run
${EPACTS_DIR}/bin/test_run_epacts.sh
You will find a series of lines in test_run_epacts.sh script commented out for each possible test.
The example phenotype (PED format) and genotype (VCF format) can be found at
${EPACTS_DIR}/share/EPACTS/
Single Variant Test
Or You can run EPACTS command yourself by running
${EPACTS_DIR}/epacts single \
--vcf ${EPACTS_DIR}/data/1000G_exome_chr20_example_softFiltered.calls.vcf.gz \
--ped ${EPACTS_DIR}/data/1000G_dummy_pheno.ped \
--min-maf 0.001 --chr 20 --pheno DISEASE --cov AGE --cov SEX --test b.score --anno \
--out out/test --run 2
The command above will perform single variant association test using a dummy case-control phenotype file and a subset of 1000 genomes exome VCF file (chr20) using score test statistic for all variants over 1% of higher MAF using 2 parallel runs.
You will see the 4 output files as the main outcome of the analysis
Output Text of All Test Statistics
The filename is out/test.single.b.score.epacts.gz and the contents will look like
$ zcat out/test.single.b.score.epacts.gz | head #CHROM BEGIN END MARKER_ID NS AC CALLRATE MAF PVALUE SCORE N.CASE N.CTRL AF.CASE AF.CTRL 20 68303 68303 20:68303_A/G_Upstream:DEFB125 266 1 1 0.0018797 NA NA NA NA NA NA 20 68319 68319 20:68319_C/A_Upstream:DEFB125 266 1.4467e-36 1 0 NA NA NA NA NA NA 20 68396 68396 20:68396_C/T_Nonsynonymous:DEFB125 266 1 1 0.0018797 NA NA NA NA NA NA 20 76635 76635 20:76635_A/T_Intron:DEFB125 266 1.534e-37 1 0 NA NA NA NA NA NA 20 76689 76689 20:76689_T/C_Synonymous:DEFB125 266 0 1 0 NA NA NA NA NA NA 20 76690 76690 20:76690_T/C_Nonsynonymous:DEFB125 266 1 1 0.0018797 NA NA NA NA NA NA 20 76700 76700 20:76700_G/A_Nonsynonymous:DEFB125 266 0 1 0 NA NA NA NA NA NA 20 76726 76726 20:76726_C/G_Nonsynonymous:DEFB125 266 0 1 0 NA NA NA NA NA NA 20 76771 76771 20:76771_C/T_Nonsynonymous:DEFB125 266 3 1 0.0056391 0.68484 0.40587 145 121 0.013793 0.0082645
Output Text of Top Associations
Same type of file but containing top 5,000 association will be stored at out/test.epacts.top5000
$ head out/test.single.b.score.epacts.top5000 #CHROM BEGIN END MARKER_ID NS AC CALLRATE MAF PVALUE SCORE N.CASE N.CTRL AF.CASE AF.CTRL 20 1610894 1610894 20:1610894_G/A_Synonymous:SIRPG 266 138.64 1 0.26061 6.9939e-05 3.9765 145 121 0.65177 0.36476 20 4162411 4162411 20:4162411_T/C_Intron:SMOX 266 204 1 0.38346 0.00055583 -3.4523 145 121 0.62759 0.93388 20 34061918 34061918 20:34061918_T/C_Intron:CEP250 266 41.815 1 0.0786 0.00095471 3.3035 145 121 0.22543 0.075436 20 4155948 4155948 20:4155948_G/A_Intron:SMOX 266 215 1 0.40414 0.0020792 -3.0787 145 121 0.68276 0.95868 20 4680251 4680251 20:4680251_A/G_Nonsynonymous:PRNP 266 186 1 0.34962 0.0025962 3.0119 145 121 0.8069 0.57025 20 36668874 36668874 20:36668874_G/A_Synonymous:RPRD1B 266 96 1 0.18045 0.003031 2.9646 145 121 0.44828 0.2562 20 36641871 36641871 20:36641871_G/A_Synonymous:TTI1 266 10 1 0.018797 0.004308 -2.8547 145 121 0.0068966 0.07438 20 1616892 1616892 20:1616892_A/G_Synonymous:SIRPG 266 144 1 0.27068 0.0051239 2.7991 145 121 0.63449 0.42975 20 25038372 25038372 20:25038372_G/A_Intron:ACSS1 266 103.3 1 0.19418 0.005748 2.7618 145 121 0.47201 0.28813
The key columns represents:
- NS : Number of phenotyped samples with non-missing genotypes
- AC : Total Non-reference Allele Count
- CALLRATE : Fraction of non-missing genotypes.
- MAF : Minor allele frequencies
- PVALUE : P-value of single variant test
- AF.CASE : Non-reference allele frequencies for cases
- AF.CTRL : Non-reference allele frequencies for controls
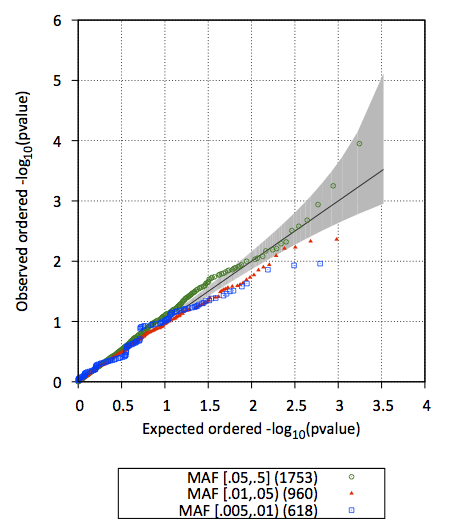
Q-Q plot of test statistics (stratified by MAF)
The file out/test.b.score.epacts.qq.pdf will be generated as shown below
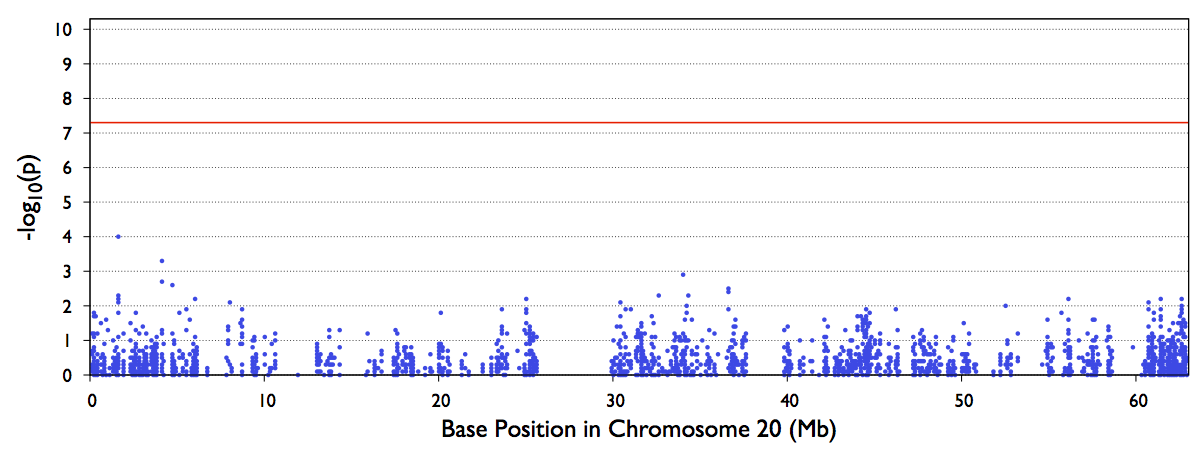
Manhattan Plot of Test Statistics
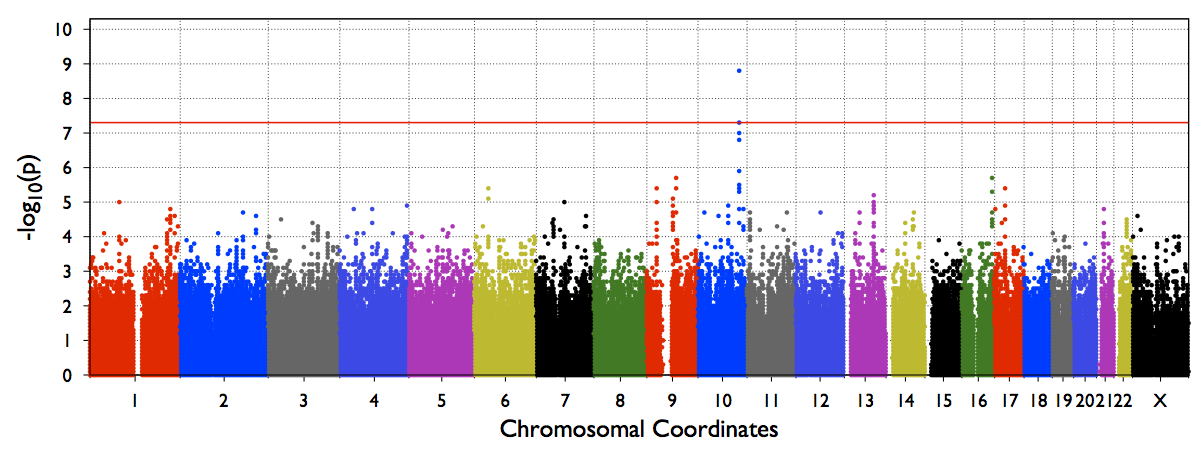
The file out/test.b.score.epacts.mh.pdf will be generated for chr20 only.
An example Genome-wide manhattan plot (from a genome-wide run) will look like below
Gene-wise or group-wise burden test
Gene-wise or group-wise burden test requires two steps. First, 'group' file containing the list of markers per group needs to be generated. Second, group-wise burden test needs to be run
Creating marker group file
The marker group file has the following format
[GROUP_ID] [MARKER_ID_1] [MARKER_ID_2] .... [MARKER_ID_N]
where
- [GROUP_ID] is a string representing the group (e.g. gene name)
- [MARKER_ID_K] is a marker key as a format of [CHROM]:[POS]_[REF]/[ALT] (NOTE THAT THIS IS DIFFERENT FROM TYPICAL VCF MARKER ID field)
Note that [MARKER_ID_K] has to be sorted by increasing order of genomic coordinate
In order to create gene-level group file from typically formatted VCF file, one may use the following utility
${EPACTS_DIR}/epacts make-group --vcf [input-vcf] --out [output-group-file] --format [epacts, annovar, chaos or gatk] --nonsyn
The above command create a file [output-group-file] containing a list of missense and nonsense variants per each gene. To incorporate different types of functional annotations, use --type option as follows
${EPACTS_DIR}/epacts make-group --vcf [input-vcf] --out [output-group-file] --format [epacts, annovar, chaos or gatk] --type [function_type_1] --type [function_type_2] ...
Type 'epacts makegroup -man' for the detailed documentation
Annotating VCF file using EPACTS
If the VCF is not annotated, 'epacts makegroup' cannot be used. In order to annotate VCF, one can use the example VCF using ANNOVAR as follows:
${EPACTS_DIR}/epacts anno \
--in ${EPACTS_DIR}/data/1000G_exome_chr20_example_softFiltered.calls.vcf.gz \
--out ${EPACTS_DIR}/data/1000G_exome_chr20_example_softFiltered.calls.anno.vcf.gz
The epacts anno script will add "ANNO=[function]:[genename]" entry into the INFO field based on gencodeV7 (default) or refGene database.
It is important to check whether the VCF file is already annotated or not in order to avoid no or redundant annotation.
Running Groupwise Test
To perform a groupwise burden test on the example VCF (annotated as above), run the following command
${EPACTS_DIR}/epacts group --vcf ${EPACTS_DIR}/data/1000G_exome_chr20_example_softFiltered.calls.anno.vcf.gz \
--groupf ${EPACTS_DIR}/data/1000G_exome_chr20_example_softFiltered.calls.anno.grp --out out/test.gene.skat \
--ped ${EPACTS_DIR}/data/1000G_dummy_pheno.ped --maxAF 0.05 \
--chr 20 --pheno QT --cov AGE --cov SEX --test skat --skat-o --run 2
Example Output
$ head out/test.gene.skat.epacts.top5000 #CHROM BEGIN END MARKER_ID NS FRAC_WITH_RARE NUM_ALL_VARS NUM_PASS_VARS NUM_SING_VARS PVALUE STATRHO 20 62607037 62608720 20:62607037-62608720_SAMD10 266 0.14662 9 5 1 0.0020064 1 20 2816211 2820493 20:2816211-2820493_FAM113A 266 0.011278 12 2 1 0.0032542 0 20 47245987 47361692 20:47245987-47361692_PREX1 266 0.1391 54 9 6 0.0054849 1 20 34761734 34810279 20:34761734-34810279_EPB41L1 266 0.071429 14 7 5 0.0068492 0.2 20 61340671 61391602 20:61340671-61391602_NTSR1 266 0.11278 24 9 3 0.011063 1 20 48561952 48568644 20:48561952-48568644_RNF114 266 0.011278 4 2 1 0.015175 0.2 20 60962895 60963559 20:60962895-60963559_RPS21 266 0.06015 6 3 2 0.016409 0 20 55904961 55917801 20:55904961-55917801_SPO11 266 0.011278 11 3 3 0.018031 0
The key columns represents:
- NS : Number of phenotyped samples with non-missing genotypes
- FRAC_WITH_RARE : Fraction of individual carrying rare variants below --max-maf (default : 0.05) threshold.
- NUM_ALL_VARS : Number of all variants defining the group.
- NUM_PASS_VARS : Number of variants passing the --min-maf, --min-mac, --max-maf, --min-callrate thresholds
- NUM_SING_VARS : Number of singletons among variants in NUM_PASS_VARS
- PVALUE : P-value of burden tests
- Other columns are test specific auxiliary columns. For example, in the VT test, the optimal MAF threshold is recorded as an auxiliary output column.
Specialized Instruction for EMMAX tests
EMMAX (Efficient Mixed Model Association eXpedited - Kang et al (2010) Nat Genet 42:348-54) is an efficient implementation of mixed model association accounting for sample structure including population structure and hidden relatedness. Currently EPACTS supports EMMAX association mapping in single variant test and CMC-like burden tests.
Because EMMAX is based on linear model, the method fits better to quantiative traits than binary traits. However, p-values for binary traits are expected to be valid in the spirit of Armitage trend test, although the estimated effect size may not be precise.
In order to run EMMAX analysis from sequence-based genotypes. We recommend running EPACTS multiple times using the following procedure.
Single Variant EMMAX Association Analysis
- Creating Kinship Matrix : From VCF, we recommend to set a MAF (e.g. 0.01) and call rate (e.g. 0.95) threshold to select high-quality markers to generate kinship matrix as follows.
${EPACTS_DIR}/epacts make-kin \
--vcf [input.vcf.gz] --ped [input.ped (Optional)] --min-maf 0.01 --minCallRate 0.95 \
--sepchr (if VCF is separated by chromosome) --out [outprefix.kinf] --run [# of parallel jobs]
If you provide [input.ped] file, then it will calculate the subset the individuals contained in the PED file.
The procedure above will create a file [outprefix.kinf] after splitting and merging the genomes into multiple pieces. If only a certain subset of SNPs needs to be considered due to target regions, LD-pruning, or any other reasons, a VCF containing the subset of markers must be created beforehand and should be used as input VCF file.
- Perform Single Variant Association : From VCF and PED, we recommend to use less stringent MAF threshold (e.g. 0.001) and call rate (e.g. 0.50) to perform single variant association
${EPACTS_DIR}/epacts single \
--vcf [input.vcf.gz] --ped [input.ped] --min-maf 0.001 --kin [outputprefix.kinf] \
--sepchr --pheno [PHENO_NAME] --cov [COV1] --cov [COV2] --test q.emmax \
--out [outprefix] --run [# of parallel jobs]
The procedure above will perform single variant association analysis compatible to other types of single variant association analyses implemented in EPACTS
Burden-style EMMAX Association Analysis
In order to run EMMAX analysis from sequence-based genotypes. We recommend running EPACTS multiple times using the following procedure.
- Creating Kinship Matrix : See 'Creating Kinship Matrix' section in #Single Variant EMMAX Association Analysis
- Create Marker Group
- By annotating the VCF and extracting missense and nonsense variants
- #Annotating VCF file using ANNOVAR - This step will be required to create marker group file
- #Creating marker group file - Assume that [group.grp] file is produced
- Or, by creating your own marker group information
- See #Creating marker group file for details
- By annotating the VCF and extracting missense and nonsense variants
- Run CMC-style burden test by
${EPACTS_DIR}/epacts group --groupf [group.grp] \
--vcf [input.vcf.gz] --ped [input.ped] --max-maf [max-MAF-for-rare-variants] \
--kin [outputprefix.kinf] --sepchr --pheno [PHENO_NAME] --cov [COV1] --cov [COV2] \
--test emmaxCMC --out [outprefix]
- Run Variable Threshold burden test by
${EPACTS_DIR}/epacts group --groupf [group.grp] \
--vcf [input.vcf.gz] --ped [input.ped] --max-maf [max-MAF-for-rare-variants] \
--kin [outputprefix.kinf] --sepchr --pheno [PHENO_NAME] --cov [COV1] --cov [COV2] \
--test emmaxVT --out [outprefix]
Preparing Your Own Input Data
VCF file for Genotypes
EPACTS support VCF files as input for association with the following requirement
- Input VCF file must be bgzipped and tabixed before running association to allow efficient random access of the file. Below is an example command to conver plain VCF into bgzipped and tabixed VCF
bgzip input.vcf ## this command will produce input.vcf.gz tabix -pvcf -f input.vcf.gz ## this command will produce input.vcf.gz.tbi
- If the VCF file is separated by chromosome, the VCF file specified in the input argument must contain the string "chr1" in the chromosome 1 file, and corresponding chromosome name for other chromosomes. Thus, the files names should be like
[prefix]chr1[suffix].vcf.gz,[prefix]chr2[suffix].vcf.gz, ...,[prefix]chr22[suffix].vcf.gz,[prefix]chrX[suffix].vcf.gz. - Sample IDs in the VCF file must be consistent to those from PED file
- Currently EPACTS only support bi-allelic variants, but it handles SNPs, INDELs, snd SVs.
- Currently, EPACTS only support VCF aligned with NCBI build 37 coordinates
- An example VCF file from 1000 genome project is below.
$ zcat example/1000G_integrated_phase1_chr20.vcf.gz | cut -f 1-10 | head -50 ##fileformat=VCFv4.1 ##INFO=<ID=LCSNP,Number=0,Type=Flag,Description="Genotype likelihood in Low coverage VCF in data integration"> ##INFO=<ID=EXSNP,Number=0,Type=Flag,Description="Genotype likelihood in Exome VCF in data integration"> ##INFO=<ID=INDEL,Number=0,Type=Flag,Description="Genotype likelihood in INDEL VCF in data integration"> ##INFO=<ID=SV,Number=0,Type=Flag,Description="Genotype likelihood in SV VCF in data integration"> ##INFO=<ID=BAVGPOST,Number=1,Type=Float,Description="Average posterior probability from beagle"> ##INFO=<ID=BRSQ,Number=1,Type=Float,Description="Genotype imputation quality estimate from beagle"> ##INFO=<ID=LDAF,Number=1,Type=Float,Description="MLE Allele Frequency Accounting for LD"> ##INFO=<ID=AVGPOST,Number=1,Type=Float,Description="Average posterior probability from MaCH/Thunder"> ##INFO=<ID=RSQ,Number=1,Type=Float,Description="Genotype imputation quality from MaCH/Thunder"> ##INFO=<ID=ERATE,Number=1,Type=Float,Description="Per-marker Mutation rate from MaCH/Thunder"> ##INFO=<ID=THETA,Number=1,Type=Float,Description="Per-marker Transition rate from MaCH/Thunder"> ##INFO=<ID=CIEND,Number=2,Type=Integer,Description="Confidence interval around END for imprecise variants"> ##INFO=<ID=CIPOS,Number=2,Type=Integer,Description="Confidence interval around POS for imprecise variants"> ##INFO=<ID=END,Number=1,Type=Integer,Description="End position of the variant described in this record"> ##INFO=<ID=HOMLEN,Number=.,Type=Integer,Description="Length of base pair identical micro-homology at event breakpoints"> ##INFO=<ID=HOMSEQ,Number=.,Type=String,Description="Sequence of base pair identical micro-homology at event breakpoints"> ##INFO=<ID=SOURCE,Number=.,Type=String,Description="Source of deletion call"> ##INFO=<ID=SVLEN,Number=1,Type=Integer,Description="Difference in length between REF and ALT alleles"> ##INFO=<ID=SVTYPE,Number=1,Type=String,Description="Type of structural variant"> ##INFO=<ID=AC,Number=.,Type=Integer,Description="Alternate Allele Count"> ##INFO=<ID=AN,Number=1,Type=Integer,Description="Total Allele Count"> ##ALT=<ID=DEL,Description="Deletion"> ##FORMAT=<ID=GT,Number=1,Type=String,Description="Genotype"> ##FORMAT=<ID=DS,Number=1,Type=Float,Description="Genotype dosage from MaCH/Thunder"> ##FORMAT=<ID=GL,Number=.,Type=Float,Description="Genotype Likelihoods"> ##FORMAT=<ID=BD,Number=1,Type=Float,Description="Genotype dosage from beagle"> #CHROM POS ID REF ALT QUAL FILTER INFO FORMAT HG00096 20 60479 . C T 100 PASS LCSNP;EXSNP;BAVGPOST=1.000;BRSQ=0.894;LDAF=0.0020;AVGPOST=0.9995;RSQ=0.8779;ERATE=0.0005;THETA=0.0008;AC=4;AN=2184 GT:DS:GL:BD 0|0:0.000:-0.19,-0.46,-2.68:0.0022 20 60522 . T TC 1588 PASS INDEL;BAVGPOST=1.000;BRSQ=0.994;LDAF=0.0116;AVGPOST=0.9980;RSQ=0.9327;ERATE=0.0004;THETA=0.0167;AC=24;AN=2184 GT:DS:GL:BD 0|0:0.000:0.00,-0.90,-9.20:0 20 60571 . C A 100 PASS LCSNP;EXSNP;BAVGPOST=0.999;BRSQ=0.813;LDAF=0.0029;AVGPOST=0.9986;RSQ=0.8085;ERATE=0.0014;THETA=0.0014;AC=5;AN=2184 GT:DS:GL:BD 0|0:0.000:-0.05,-0.96,-5.00:0.0008 20 60795 . G C 100 PASS LCSNP;EXSNP;BAVGPOST=1.000;BRSQ=0.930;LDAF=0.0006;AVGPOST=0.9996;RSQ=0.7205;ERATE=0.0003;THETA=0.0041;AC=1;AN=2184 GT:DS:GL:BD 0|0:0.000:-0.03,-1.21,-5.00:0.0001 20 60810 . G GA 127 PASS INDEL;BAVGPOST=1.000;BRSQ=0.862;LDAF=0.0013;AVGPOST=0.9987;RSQ=0.5684;ERATE=0.0004;THETA=0.0061;AC=2;AN=2184 GT:DS:GL:BD 0|0:0.000:0.00,-1.80,-18.80:0
PED file for Phenotypes and Covariates
EPACTS accepts a PED format supported by MERLIN or PLINK software to represent phenotypes. For example, the example.ped file and example.dat file can represent the phenotypes and corresponding column name (from 6th column and after).
$ head example.ped 13281 NA12344 NA12347 NA12348 1 1 94.17 66.1 13281 NA12347 0 0 1 1 109.54 44.0 13281 NA12348 0 0 2 2 119.40 46.6 1328 NA06984 0 0 1 2 87.72 39.3 1328 NA06989 0 0 2 1 100.60 41.7 1328 NA12329 NA06984 NA06989 2 1 100.85 46.4 13291 NA06986 0 0 1 2 91.94 61.9 13291 NA06995 NA07435 NA07037 1 2 104.36 57.4 13291 NA06997 NA06986 NA07045 2 2 107.53 53.1
$ cat example.dat A DISEASE T QT T AGE
EPACTS also accept a PED format with header information. The above file can be combined into one file as follows
$ head data/1000G_dummy_pheno.ped #FAM_ID IND_ID FAT_ID MOT_ID SEX DISEASE QT AGE 13281 NA12344 NA12347 NA12348 1 1 94.17 66.1 13281 NA12347 0 0 1 1 109.54 44.0 13281 NA12348 0 0 2 2 119.40 46.6 1328 NA06984 0 0 1 2 87.72 39.3 1328 NA06989 0 0 2 1 100.60 41.7 1328 NA12329 NA06984 NA06989 2 1 100.85 46.4 13291 NA06986 0 0 1 2 91.94 61.9 13291 NA06995 NA07435 NA07037 1 2 104.36 57.4 13291 NA06997 NA06986 NA07045 2 2 107.53 53.1
The column names can be used to identify the names of phenotypes and covariates in the analysis.
Frequently Asked Questions
Installation
- How should I install EPACTS?
- I am having the following error message configure: error: libR.{so,a} was not found. Please install it at http://www.r-project.org/ first. What do I have to do?
- First, you need to find out where R was installed. Try to type "locate libR.so" and see if it returns anything
- If "locate libR.so" returns you something, as explained Installation Details, try to add "LDFLAGS=-L/path/to/R/library" and rerun configure and make
- If you cannot find libR.so, you make have to recompile R with --enable-R-shlib option as described in http://cran.r-project.org/doc/manuals/R-admin.html#Installation
Input Files
- What is VCF?
- VCF refers to Variant Call Format
- See [1000 Genomes wiki page] for the detailed description of VCF format
- Should input VCF be compressed into certain format?
- Correct. EPACTS assumes that VCF file is bgzipped and tabixed already.
- See #VCF file for Genotypes for details.
- What are the additional requirements for input VCF file?
- Input VCF file used for association mapping must contain individual genotype information at 10-th or higher order columns.
- GT field must be encoded as haploid or diploid
- Bi-allelic SNPs only : Currently EPACTS may not handle multi-allelic SNPs correctly.
- If non-GT field is used, the field is considered as dosage and should be a single numeric value.
- What are the acceptable input format to encode phenotypes and covariates?
- See #PED file for Phenotypes and Covariates for the detailed information
- How should I encode binary phenotypes?
- If you encode your phenotypes into two different numeric values (e.g. 0/1 or 1/2), EPACTS will automatically recognize them as binary phenotypes and encode them into 1/2 values. Higher value will be considered as cases for case-control association
- How should I encode missing genotypes?
- The default code missing phenotypes in EPACTS are 'NA'
- One may use --missing option to specify different types of missing values
- The encoding of missing genotypes follows the VCF specificiation
- How do I match the relationship between VCF and PED input files?
- EPACTS will assume that the individual IDs in each VCF and PED file are unique, and they follow the saming convention. Thus, the individual IDs overlapping between VCF and PED files will be considered in the associations
- How the individuals with missing phenotypes are handled?
- Currently, EPACTS will automatically remove the individuals without phenotypes or covariates. If one wants to use imputed covariates to increase sample size, the PED file must contain the imputed covariate values.
- Markers with missing genotypes won't be discarded automatically. It can be explicitly discarded by --minCallRate option when performing association
Output Files
- Which output files should I be looking at?
- #Output Text of Top Associations is the key file to look at the individual top associations
- #Q-Q plot of test statistics (stratified by MAF) will be important to see the global distribution of test statistics and examine if there are apparent inflation of test statistics
- #Manhattan Plot of Test Statistics will inform us the genome-wide distribution of association signals
- #Output Text of All Test Statistics will contain the full information of test results across all units tested
- The Q-Q and Manhattan plots cannot be found. Why?
- It is probably because gnuplot 4.2 or higher is not installed in your system, or they are included but cannot be found in your ${PATH}. Please visit [GNUPLOT web page] for installation.
- How can I read the EMMAX kinship file from produced from EPACTS?
- * You can run the following command to dump your kinship matrix into a human-readable text format.
$(EPACTS_DIR)/bin/pEmmax kin-util --kinf [input.kinf] --outf [output.prefix] --dump
More questions
- If you have more questions, please contact [Hyun Min Kang].
Detailed Options
The detailed options can viewed by running the following commands
${EPACTS_DIR}/bin/epacts -man (for overall structure)
${EPACTS_DIR}/bin/epacts single -man (for single variant test)
${EPACTS_DIR}/bin/epacts group -man (for groupwise test)
${EPACTS_DIR}/bin/epacts anno -man (for annotation)
${EPACTS_DIR}/bin/epacts plot -man (for QQ and Manhattan plot)
${EPACTS_DIR}/bin/epacts zoom -man (for zoom plot)
${EPACTS_DIR}/bin/epacts meta -man (for meta-analysis)
${EPACTS_DIR}/bin/epacts make-group -man (for creating gene group)
Implementing Additional Statistical Tests
In order to add additional statistical test to EPACTS, the following procedure are recommended
- Create a file named 'single.[testname].R' for single variant test or 'gene.[testname].R' for gene-level test under ${EPACTS_DIR}/share/EPACTS/
- Test your implementation using --test [testname] option to perform sanity check and debugging
- If you want to add your test in the official in-house version, please send your code to Hyun
Below is an example of a single variant test implementation ( single.q.lm.R )
## Core functions of EPACTS to perform association
##################################################################
## SINGLE VARIANT TEST
## INPUT VARIABLES:
## n : total # of individuals
## NS : number of called samples
## AC : allele count
## MAF : minor allele frequency
## vids : indices from 1:nrow(NS) after AF/AC threshold
## genos : genotype matrix (after AF/AC threshold)
## EXPECTED OUTPUT : list(p, addcols, addnames) for each genos row
## p : p-value
## add : additional columns to add
## cname : column names for additional columns
##################################################################
## single.lm() : Use built-in lm() function to perform association
## KEY FEATURES : SIMPLE, BUT MAY BE SLOW
## GOOD SNIPPLET TO START A NEW FUNCTION
## TRAITS : QUANTITATIVE
## RETURNS : PVALUE, BETA, SEBETA, TSTAT
## MISSING VALUES : IGNORED
single.q.lm <- function() {
cname <- c("BETA","SEBETA","TSTAT") # column names for additional variables in the EPACTS output
m <- nrow(genos)
p <- rep(NA,m)
add <- matrix(NA,m,3) ## BETA, SEBETA, TSTAT
if ( m > 0 ) {
for(i in 1:m) {
r <- summary(lm(pheno~genos[i,]+cov-1))$coefficients[1,] # run simple linear regression
p[i] <- r[4] # store p-value to p[i]
add[i,] <- r[1:3] # store additional variables to add[i,]
}
}
return(list(p=p,add=add,cname=cname))
}
As described in the comment, you may assume that the following variables are available for use for testing association across m markers
- n (scalar) : total number of individuals
- NS (M * 1 vector) : Number of called samples for each marker
- AC (M * 1 vector) : Non-reference allele count for each marker
- MAF (M * 1 vector) : Minor allele frequency
- vids (m * 1 vector) : indices of markers passing the inclusion criteria (e.g. MAF threshold) among 1:M
- genos (m * n matrix) : genotype matrix as a input for association test
The output variables to generate is as follows
- p (m * 1 vector) : p-value matrix as output
- add (m * c matrix) : additional columns as output of test (such as SCORE, BETA, etc)
- cname (c * 1 vector) : column names of add
In the output files, the following columns will be displayed
- MARKER : Marker ID
- NS : Number of called samples
- AC : Non-ref allele count
- CALLRATE : Call rate = NS/n
- MAF : Minor allele frequency
- PVALUE : P-values
- Additional columns specified by return values 'add'
Below is an example of a gene-lvel variant test implementation ( single.q.lm.R )
##################################################################
## GENE-LEVEL BURDEN TEST
## INPUT VARIABLES:
## n : total # of individuals
## genos : genotype matrix for each gene
## NS : number of called samples for each marker
## AC : allele count for each marker
## MAC : minor allele count for each marker
## MAF : minor allele frequency
## vids : indices from 1:n after AF/AC threshold
## EXPECTED OUTPUT : list(p, addcols, addnames) for each genos row
## p : p-value
## add : additional column values
## cname : additional column names
##################################################################
## gene.q.reverse() : Reverse logistic regression
## KEY FEATURES : 0/1 collapsing variable ~ rare variants
## TRAITS : QUANTITATIVE (GAUSSIAN)
## RETURNS : PVALUE, BETA, SEBETA, ZSTAT
## MISSING VALUE : IMPUTED AS MAJOR ALLELES
gene.q.reverse <- function() {
cname <- c("BETA","SEBETA","ZSTAT")
m <- nrow(genos)
if ( m > 0 ) {
g <- as.double(colSums(genos,na.rm=T) > 0)
sg <- sum(g)
if ( ( sg > 0 ) && ( sg < n ) ) {
r <- glm(g~pheno+cov-1,family=binomial)
if ( ( r$converged ) && ( ! r$boundary ) ) {
return(list(p=summary(r)$coefficients[1,4],
add=summary(r)$coefficients[1,1:3],
cname=cname))
}
}
}
return(list(p=NA,add=rep(NA,3),cname=cname))
}
Similar to gene-level test, you may assume the following variables exist for testing A SINGLE GENE. Note that M is the number of markers spanning the gene region
- n (scalar) : total number of individuals
- NS (M * 1 vector) : Number of called samples for each marker
- AC (M * 1 vector) : Non-reference allele count for each marker
- MAC (M * 1 vector) : Minor allele count
- MAF (M * 1 vector) : Minor allele frequency
- vids (m * 1 vector) : indices of markers passing the inclusion criteria (e.g. MAF threshold) among 1:M
- genos (m * n matrix) : genotype matrix as a input for association test
The output variables to generate is as follows
- p (scalar) : p-value matrix as output
- add (c * 1 vector) : additional columns as output of test (such as SCORE, BETA, etc)
- cname (c * 1 vector) : column names of add
In the output files, the following columns will be displayed
- MARKER : Marker ID
- NS : Number of called samples
- MAF_BURDEN : MAF of 0/1 collapsing variables (existence of rare variants)
- NUM_ALL_VARS : Number of all variants within the gene
- NUM_RARE_VARS : Number of rare variants below the max-MAF threshold
- NUM_SING_VARS : Number of singleton variants
- PVALUE : P-value from the test
- Additional columns specified by return values 'add'
Full ChangeLog
- July 10th, 2014 : EPACTS v3.2.6 release
- Minor bug fix in epacts-make-kin
- March 11th, 2014 : EPACTS v3.2.5 release
- EMMAX-SKAT is implemented with major bug fix
- November 21th, 2013 : EPACTS v3.2.4 release
- Fixed a number of minor bugs
- Some known bugs still exist
- SKAT-O Lambda eigenvalue error. This happenes in a particular context but haven't nailed down a way to prevent it yet.
- EMMAX has case and control frequency flipped.
- EMMAX test has a silly known bug with case / ctrl frequency is flipped
- March 25th, 2013 : EPACTS v3.2.3 release
- Relaxed the checking of low-rank matrix in SKAT tests (to avoid unncessary skipping of genes)
- March 13th, 2013 : EPACTS v3.2.2 release
- Fixed an error which occasionally report mismatches in the number of samples
- March 9th, 2013 : EPACTS v3.2.1 release
- Fixed errors in loading the dynamic library
- Fixed errors in SKAT-O (thanks to Anubha Mahajan and Jason Flannick)
- Fixed bugs in emmax-CMC
- Added emmax-SKAT (contributed by Seunngeun Lee)
- And additional minor bug fixes
- February 28th, 2013 : EPACTS v3.2.0 release
- R package installation bug (for some users) was fixed
- A bug in the MAF error for high frequency variants (AF>0.25) was now fixed
- SKAT version is updated to 0.81
- --bprange option is added to allow testing for small region size
- Additional minor bug fixes
- December 4th, 2012 : EPACTS v3.1.0 release
- Removed dependency on libR.so
- Additional minor bug fixes
- --bprange option is added to allow testing for small region size
- November 25th, 2012 : EPACTS v3.0.0 release
- Restructured with source code release (with autoconf / automake / libtools)
- Added zoom plot feature
- FRAC_BURDEN keyword was replace to FRAC_WITH_RARE for groupwise testing
- October 26th, 2012 : EPACTS v2.2.0-beta is released with the following updates
- Added --max-mac option
- Fixed Firth's bias-corrected test (by Clement Ma)
- Added more informative warning messages when index files do not exist
- Fixed the bug in the epacts-plot in plotting ties
- Fixed errors in the MAF estimates per case and control
- Fixed bug in --minRSQ option
- September 28, 2012 : EPACTS v2.11-beta is released with the following updates
- Counts and allele frequencies for case/control added for binary tests
- --max-maf parameter is added
- Fixed EMMAX error in MAF in the output
- More informative error messages
- September 27, 2012 : EPACTS v2.1-beta is released with the following updates
- EMMAX interface is changed. --kinOnly option is related with a new command make-kin
- SKAT-O is upgraded to version 0.77 with additional configurable parameter settings
- Some parameter names are renamed (e.g. --min-maf, --min-mac)
- Many minor bugs are fixed
- Jul 6, 2012 : EPACTS v2.01-beta is released with the following updates
- SKAT-O is upgraded to version 0.76
- Fixed minor bugs in option names (Thanks to Xueling Sim)
- Jul 3, 2012 : EPACTS v2.0-beta is released with the following updates
- Major restructuring of the software
- Annotation software is switched with built-in application
- Addition of SKAT-O and EMMAX burden test
- Minor bug fixes
- Apr 8, 2012 : EPACTS v1.2-alpha is released with the following updates, in addition to the following updates
- EMMAX bug in handling covariates was fixed
- Variable Threshold Test is added
- Variable Threshold Test with genomic score (e.g. GERP or PhyloP) is added.
- Apr 4, 2012 : EPACTS v1.1-alpha is released with the following updates, in addition to minor updates
- EMMAX burden test (Hyun Min Kang)
- Likelihood ratio test (Clement Ma)
- Updated version of Firth bias-corrected likelihood ratio test (Clement Ma)
- Updated version of EMMAX single variant test (Hyun Min Kang)
- Mar 29, 2012 : EPACTS v1.0-alpha is released